| Revision as of 16:14, 14 August 2014 editJohn of Reading (talk | contribs)Autopatrolled, Extended confirmed users, Pending changes reviewers767,601 editsm →Discovery: Typo fixing, replaced: the the → the using AWB← Previous edit | Latest revision as of 20:24, 10 January 2025 edit undoOnel5969 (talk | contribs)Autopatrolled, Extended confirmed users, Page movers, New page reviewers, Pending changes reviewers, Rollbackers937,500 editsm Disambiguating links to Papuans (link changed to Indigenous people of New Guinea; link changed to Indigenous people of New Guinea) using DisamAssist. | ||
| Line 1: | Line 1: | ||
| {{short description|Asian archaic human}} | |||
| {{Redirect3|Woman X|For other uses, see ] or ]}} | |||
| {{good article}} | |||
| {{Use dmy dates|date=November 2013}} | |||
| {{Use dmy dates|date=September 2020}} | |||
| '''Denisovans''' or '''Denisova hominins''' {{IPAc-en|d|ə|ˈ|n|iː|s|ə|v|ə}} are a ]-era species of the genus '']'' or subspecies of '']''. In {{Nowrap|March 2010}}, scientists announced the discovery of a finger bone fragment of a juvenile female who lived about 41,000 years ago, found in the remote ] in the ] in Siberia, a cave which has also been inhabited by ]s and ].<ref>{{cite web |author=David Leveille |title=Scientists Map An Extinct Denisovan Girl's Genome |url=http://www.theworld.org/2012/08/scientists-map-an-extinct-denisovan-girls-genome/ |publisher=], |date=31 August 2012 |accessdate=31 August 2012}}</ref><ref name=washingtonpost>{{Citation |url=http://www.washingtonpost.com/wp-dyn/content/article/2010/03/24/AR2010032401926_pf.html |title=DNA from bone shows new human forerunner, and raises array of questions |newspaper=] |date=25 March 2010 |first=David |last=Brown }}</ref><ref name="Pääbo et al.">{{Citation |first=Johannes |last=Krause |first2=Qiaomei |last2=Fu |first3=Jeffrey M. |last3=Good |first4=Bence |last4=Viola |first5=Michael V. |last5=Shunkov |first6=Anatoli P. |last6=Derevianko |lastauthoramp=yes |first7=Svante |last7=Pääbo |year=2010 |title=The complete mitochondrial DNA genome of an unknown hominin from southern Siberia |journal=] |pmid=20336068 |volume=464 |issue=7290 |pages=894–897 |doi=10.1038/nature08976 }}</ref> Two teeth and a toe bone belonging to different members of the same population have since been reported. | |||
| ]]] | |||
| The '''Denisovans''' or '''Denisova hominins''' ({{IPAc-en|d|ə|ˈ|n|iː|s|ə|v|ə}} {{respell|də|NEE|sə|və}}) are an ] ] or ] of ] that ranged across Asia during the ] and ], and lived, based on current evidence, from 285 to 25 thousand years ago.<ref name="NYT-20240302">{{cite news |last=Zimmer |first=Carl |authorlink=Carl Zimmer |title=On the Trail of the Denisovans - DNA has shown that the extinct humans thrived around the world, from chilly Siberia to high-altitude Tibet — perhaps even in the Pacific islands. |url=https://www.nytimes.com/2024/03/02/science/denisovan-neanderthal-dna.html |date=2 March 2024 |work=] |url-status=live |archiveurl=https://archive.today/20240302121615/https://www.nytimes.com/2024/03/02/science/denisovan-neanderthal-dna.html |archivedate=2 March 2024 |accessdate=2 March 2024 }}</ref> Denisovans are known from few physical remains; consequently, most of what is known about them comes from ] evidence. No formal species name has been established pending more complete fossil material. | |||
| Analysis of the ] (mtDNA) of the finger bone showed it to be genetically distinct from the mtDNAs of Neanderthals and modern humans.<ref name="The Scientist"/> Subsequent study of the ] from this specimen suggests that this group shares a common origin with Neanderthals, that they ranged from Siberia to Southeast Asia, and that they lived among and interbred with the ancestors of some present-day modern humans, with about 3% to 5% of the DNA of ] and ] deriving from Denisovans.<ref name=nytimes>{{cite news |title=Denisovans Were Neanderthals' Cousins, DNA Analysis Reveals |author=] |url=http://www.nytimes.com/2010/12/23/science/23ancestor.html?hp |newspaper=NYTimes.com |date=22 December 2010 |accessdate=22 December 2010}}</ref><ref name="Callaway.">{{Citation |url=http://www.nature.com/news/2011/110922/full/news.2011.551.html |title=First Aboriginal genome sequenced |publisher=] |date=22 September 2011 |first=Ewen |last=Callaway |doi=10.1038/news.2011.551}}</ref><ref>"About 3% to 5% of the DNA of people from Melanesia (islands in the southwest Pacific Ocean), Australia and New Guinea as well as aboriginal people from the Philippines comes from the Denisovans." --Elizabeth Landau's interview of Svante Paabo, accessdate=2013-12-10</ref> Other ethnicities, such as the ], ], the ] of ], ], and ]-speaking peoples may be included in this category as well.{{citation needed|date=May 2014}} A comparison with the genome of a Neanderthal from the same cave revealed significant local interbreeding, with local Neanderthal DNA representing 17% of the Denisovan genome, while evidence was also detected of interbreeding with an as yet unidentified ancient human lineage.<ref name=Pennisi>{{Citation |url=http://www.sciencemag.org/content/340/6134/799.summary |first=Elizabeth |last=Pennisi |year=2013 |title=More Genomes from Denisova Cave Show Mixing of Early Human Groups |journal=] |volume=340 |page=799 |doi=10.1126/science.340.6134.799 |issue=6134}}</ref> Similar analysis of a toe bone discovered in 2011 is underway,<ref name=newscientist>{{Citation |url=http://www.newscientist.com/article/mg21128254.000-stone-age-toe-could-redraw-human-family-tree.html |title=Stone Age toe could redraw human family tree |journal=] |date=13 August 2011 |first=Colin |last=Barras }}</ref> while analysis of DNA from two teeth found in different layers than the finger bone revealed an unexpected degree of mtDNA divergence among Denisovans.<ref name=Pennisi /> In 2013, mitochondrial DNA from a 400,000-year-old hominin femur bone from Spain, which had been seen as either Neanderthal or '']'', was found to be closer to Denisovan mtDNA than to Neanderthal mtDNA.<ref name="NAT-20131204">{{cite journal |last=Callaway |first=Ewan |title=Hominin DNA baffles experts|url=http://www.nature.com/news/hominin-dna-baffles-experts-1.14294 |date=5 December 2013|journal=] |volume=504 |pages=16–17 |doi=10.1038/504016a |accessdate=4 December 2013 }}</ref> | |||
| The first identification of a Denisovan individual occurred in 2010, based on ] (mtDNA) extracted from a ] female finger bone excavated from the Siberian ] in the ] in 2008.<ref name="Pääbo et al."/> ] indicates close affinities with ]s. The cave was also periodically inhabited by Neanderthals, but it is unclear whether Neanderthals and Denisovans ever cohabited in the cave. Additional specimens from Denisova Cave were subsequently identified, as was a single specimen from the ] on the ], and Cobra Cave in the ] of Laos. DNA evidence suggests they had dark skin, eyes, and hair, and had a Neanderthal-like build and facial features. However, they had larger ] which are reminiscent of ] to ] archaic humans and ]s. | |||
| ==Discovery== | |||
| ] | |||
| The ] is located in southwestern ], in the ] near the border with ] and ]. It is named after Denis, a Russian hermit who lived there in the 18th century. The cave was originally explored in the 1970s by Russian ] Nikolai Ovodov, who was looking for remains of cave bears.{{citation needed|date=April 2014}} In 2008, Michael Shunkov from the ] and other Russian ] from the Institute of Archaeology and Ethnology of ] investigated the cave. They found the finger bone of a juvenile ], dubbed the "X woman" (referring to the maternal descent of mitochondrial DNA<ref name="The Guardian"/>) or the Denisova hominin. Artifacts, including a bracelet, excavated in the cave at the same level were ] to around 40,000 ]. Excavations have since revealed human artifacts showing an intermittent presence going back 125,000 years.<ref>Marshall, Michael (April 2014), "Mystery Relations" in ''New Scientist'' (5, April 2014)</ref> | |||
| Denisovans apparently ] with modern humans, with a high percentage (roughly 5%) occurring in ], ], and ]. This distribution suggests that there were Denisovan populations across Asia. There is also evidence of interbreeding with the Altai Neanderthal population, with about 17% of the Denisovan genome from Denisova Cave deriving from them. A first-generation hybrid nicknamed "]" was discovered with a Denisovan father and a Neanderthal mother. Additionally, 4% of the Denisovan genome comes from an unknown archaic human species, which diverged from modern humans over one million years ago. | |||
| A team of scientists led by ] and ] from the ] in Leipzig, Germany, ] extracted from the fragment. The cool climate of the Denisova Cave preserved the DNA.<ref name="Pääbo et al." /> The average annual temperature of the cave remains at 0 °C, which has contributed to the preservation of archaic DNA among the remains discovered.<ref name="NYT-01302012">{{cite news |last=Mitchell |first=Alanna|title=DNA Turning Human Story Into a Tell-All|url=http://www.nytimes.com/2012/01/31/science/gains-in-dna-are-speeding-research-into-human-origins.html|date=30 January 2012 |publisher=] |accessdate=31 January 2012 }}</ref> The analysis indicated that modern humans, ], and the Denisova hominin last shared a common ancestor around {{Nowrap|1 million}} years ago.<ref name="The Scientist">{{Citation |url=http://www.the-scientist.com/blog/display/57254/#ixzz0j820ioz1 |title=New hominin found via mtDNA |work=The Scientist |date=24 March 2010 |first=Alla |last=Katsnelson }}</ref> | |||
| == Taxonomy == | |||
| The mtDNA analysis further suggested this new hominin species was the result of an earlier migration ], distinct from the later out-of-Africa migrations associated with modern humans, but also distinct from the earlier African exodus of '']''.<ref name="The Scientist"/> Pääbo noted the existence of this distant branch creates a much more complex picture of humankind during the ].<ref name="The Guardian">{{Citation |url=http://www.guardian.co.uk/science/2010/mar/24/new-human-species-siberia |title=New species of human ancestor found in Siberia |work=The Guardian |date=24 March 2010 |first=Ian |last=Sample }}</ref> This work shows that the Denisovans were actually a sister group to the Neanderthals,<ref>''Nature'' Vol 468, p.1053</ref> branching off from the human lineage 600,000 years ago, and diverging from Neanderthals, probably in the Middle East, 200,000 years later.<ref>Marchall, Michael (April), op cit p.36</ref> | |||
| Denisovans may represent a new species of '']'' or an archaic subspecies of '']'' (modern humans), but there are too few fossils to erect a proper ]. Proactively proposed species names for Denisovans are ''H. denisova''<ref>{{Cite book|url=https://books.google.com/books?id=sHyFCwAAQBAJ&q=%22homo+denisova%22&pg=PA324|title=Exploring Human Biology in the Laboratory|last1=Douglas|first1=M. M.|last2=Douglas|first2=J. M.|year=2016|publisher=Morton Publishing Company|isbn=9781617313905|page=324|access-date=25 October 2020|archive-date=1 September 2024|archive-url=https://web.archive.org/web/20240901161942/https://books.google.com/books?id=sHyFCwAAQBAJ&q=%22homo+denisova%22&pg=PA324#v=snippet&q=%22homo%20denisova%22&f=false|url-status=live}}</ref> or ''H. altaiensis''.<ref>{{Cite journal|last1=Zubova |first1=A. |last2=Chikisheva |first2=T. |last3=Shunkov |first3=M. V. |year=2017 |title=The Morphology of Permanent Molars from the Paleolithic Layers of Denisova Cave |url=https://www.researchgate.net/publication/315970345 |journal=]|volume=45|pages=121–134|doi=10.17746/1563-0110.2017.45.1.121-134 |doi-access=free}}</ref> Chinese researchers suggest the Denisovans were members of '']'', and the idea has been supported by the palaeontologist ].<ref>{{Cite news |last1=McKie |first1=Robin |date=2024-03-30 |title=Scientists link elusive human group to 150,000-year-old Chinese 'dragon man' |url=https://www.theguardian.com/science/2024/mar/30/scientists-link-elusive-human-group-to-150000-year-old-chinese-dragon-man |access-date=2024-03-31 |work=The Observer |language=en-GB |issn=0029-7712 |archive-date=1 September 2024 |archive-url=https://web.archive.org/web/20240901161934/https://www.theguardian.com/science/2024/mar/30/scientists-link-elusive-human-group-to-150000-year-old-chinese-dragon-man |url-status=live }}</ref> In 2024, paleoanthropologists Christopher Bae and Xiujie Wu designated the Xujiayao hominin fossils as the holotype of the species '']'', and suggested sinking Denisovans into this species.<ref name=Bae2024>{{Cite book|last=Bae |first=C. J. |chapter=The "Muddle in the Middle" (~400 ka–~100 ka) |title=The Paleoanthropology of Eastern Asia |year=2024 |pages=95–131 |publisher=University of Hawai‘i Press |doi=10.1515/9780824898106-007 |isbn=9780824898106 }}</ref> | |||
| === Discovery === | |||
| Later in 2010, a second paper from the Svante Pääbo group reported the prior discovery, in 2000, of a third upper molar from a young adult, dating from about the same time (the finger was from level 11 in the cave sequence, the tooth from level 11.1). The tooth differed in several aspects from those of Neanderthals, while having archaic characteristics similar to the teeth of ''Homo erectus''. They performed mitochondrial DNA analysis on the tooth and found it to have a sequence different from but similar to that of the finger bone, indicating a divergence time about 7,500 years before, and suggesting it belonged to a different individual from the same population.<ref name="Reich et al.">{{Citation |first1=David |last1=Reich | |||
| {{Location map many| Asia | |||
| |first2=Richard E. |last2=Green | |||
| | relief = yes | |||
| |first3=Martin |last3=Kircher | |||
| | caption = Locations of paleoarchaeological finds linked to Denisovans: Denisova Cave (blue) in the ] of ]; ] (yellow) on the ]; and Tam Ngu Hao 2 Cave (grey) in northern ] | |||
| |first4=Johannes |last4=Krause | |||
| | coordinates1 = {{coord|51|23|51|N|84|40|34|E}} | |||
| |first5=Nick |last5=Patterson | |||
| | label1 = Denisova Cave | |||
| |first6=Eric Y. |last6=Durand | |||
| | mark1 = blue pog.svg | |||
| |first7=Bence |last7=Viola | |||
| | coordinates2 = {{coord|35|26|53|N|102|34|17|E}} | |||
| |first8=Adrian W. |last8=Briggs | |||
| | label2 = Baishiya Karst Cave | |||
| |first9=Udo |last9=Stenzel | |||
| | mark2 = Yellow pog.svg | |||
| |display-authors=9 |lastauthoramp=yes |year=2010 |title=Genetic history of an archaic hominin group from Denisova Cave in Siberia |journal=] |volume=468 |issue=7327 |pages=1053–1060 |doi=10.1038/nature09710 |pmid=21179161}}</ref> | |||
| | coordinates3 = {{coord|20|12|41.5|N|103|24|32.2|E}} | |||
| | label3 = Tam Ngu Hao 2 Cave | |||
| | mark3 = Black pog.svg | |||
| }} | |||
| ], where the first reported Denisovans were found]] | |||
| ] is located in ], Russia, in south-central ], on the western edges of the ]. It is named after Denis (Dyonisiy), a Russian ] ] who lived there in the 18th century. The cave was first inspected for fossils in the 1970s by ] ] Nikolai Ovodov, who was looking for remains of ]s.<ref>{{cite journal |title= A 33,000-Year-Old Incipient Dog from the Altai Mountains of Siberia: Evidence of the Earliest Domestication Disrupted by the Last Glacial Maximum|year=2011 |doi=10.1371/journal.pone.0022821 |volume=6 |issue=7 |journal=] |pages=e22821 |pmid=21829526 |pmc=3145761 |last1=Ovodov |first1=N. D. |last2 =Crockford |first2=S. J. |last3=Kuzmin |first3=Y. V. |last4=Higham |first4=T. F. |last5=Hodgins |first5=G. W. |last6=van der Plicht |first6=J. |display-authors=4 |bibcode= 2011PLoSO...622821O |doi-access=free}}</ref> | |||
| In 2008, ] from the ] and other Russian ] from the Institute of Archaeology and Ethnography of the ] in ] ] investigated the cave and found the finger bone of a juvenile female ] originally dated to 50–30,000 years ago.<ref name="Pääbo et al."/><ref>{{cite book|first=D. |last=Reich |author-link=David Reich (geneticist) |title=Who We Are and How We Got Here |publisher=] |year=2018 |page=53 |isbn=978-0-19-882125-0}}</ref> The estimate has changed to 76,200–51,600 years ago.<ref name= Douka/> The specimen was originally named X-woman because ] ] (mtDNA) extracted from the bone demonstrated it to belong to a novel ancient hominin, genetically distinct both from contemporary modern humans and from ]s.<ref name="Pääbo et al.">{{cite journal|first1=J. |last1=Krause |first2=Q.|last2=Fu |first3=J. M. |last3=Good |first4=B. |last4=Viola |first5=M. V. |last5=Shunkov |first6=A. P. |last6=Derevianko |name-list-style=amp |first7=S. |last7=Pääbo |author7-link=Svante Pääbo |display-authors=4 |year=2010 |title=The complete mitochondrial DNA genome of an unknown hominin from southern Siberia |volume=464 |issue=7290 |pages=894–897 |doi=10.1038/nature08976 |bibcode=2010Natur.464..894K |doi-access=free |journal=] |language=en |pmid=20336068 |s2cid=4415601 |issn=1476-4687|pmc=10152974 }}</ref> | |||
| In 2011, a toe bone was discovered in the cave, in layer 11, and therefore contemporary with the finger bone. Preliminary characterization of the bone's mitochondrial DNA suggests it belonged to a Neanderthal, not a Denisovan.<ref name="AAAS-Gibbons">{{Cite journal | |||
| | last = Gibbons | first = Ann | title = Who Were the Denisovans? | |||
| | journal = Science | volume = 333 |date=August 2011 | pages = 1084–87 | |||
| | url = http://genetics.med.harvard.edu/reich/Reich_Lab/Press_files/2011_Science-Denisova-1084-7.pdf | |||
| | accessdate = January 2012 | doi=10.1126/science.333.6046.1084 | |||
| }}</ref> The cave also contains stone tools and bone artifacts made by modern humans, and Pääbo commented: "The one place where we are sure all three human forms have lived at one time or another is here in Denisova Cave."<ref name="AAAS-Gibbons" /> | |||
| In 2019, Greek archaeologist ] and colleagues ] specimens from Denisova Cave, and estimated that Denisova 2 (the oldest specimen) lived 195,000–122,700 years ago.<ref name=Douka>{{cite journal |last1=Douka |first1=K. |title=Age estimates for hominin fossils and the onset of the Upper Palaeolithic at Denisova Cave |journal=] |date=2019 |volume=565 |issue=7741 |pages=640–644 |doi=10.1038/s41586-018-0870-z |pmid=30700871 |url=https://ro.uow.edu.au/cgi/viewcontent.cgi?article=1559&context=smhpapers1 |bibcode=2019Natur.565..640D |s2cid=59525455 |access-date=7 December 2019 |archive-date=6 May 2020 |archive-url=https://web.archive.org/web/20200506140551/https://ro.uow.edu.au/cgi/viewcontent.cgi?article=1559&context=smhpapers1 |url-status=live}}</ref> Older Denisovan DNA collected from sediments in the East Chamber dates to 217,000 years ago. Based on ] also discovered in the cave, hominin occupation (most likely by Denisovans) began 287±41 or 203±14 ]. Neanderthals were also present 193±12 ka and 97±11 ka, possibly concurrently with Denisovans.<ref name=Jacobs2019/> | |||
| ==Anatomy== | |||
| Little is known of the precise anatomical features of the Denisovans since the only physical remains discovered thus far are the ], two teeth from which genetic material has been gathered and a ]. The single finger bone is unusually broad and robust, well outside the variation seen in modern people. Surprisingly, it belonged to a female, indicating the Denisovans were extremely robust, perhaps similar in build to the Neanderthals. The tooth that has been characterized shares no derived morphological features with Neanderthal or modern humans.<ref name="Reich et al."/> An initial morphological characterization of the toe bone led to the suggestion that it may have belonged to a Neanderthal-Denisovan hybrid individual, although a critic suggested the morphology was inconclusive. This toe bone is currently undergoing DNA analysis by Pääbo.<ref name="newscientist"/> | |||
| === Specimens === | |||
| Some older finds may or may not belong to the Denisovan line. These includes the skulls from ] and ], and a number of more fragmentary remains from Asia. Asia is not well mapped with regard to human evolution, and the above finds may represent a group of "Asian Neanderthals". | |||
| {{redirect|Woman X||X woman (disambiguation)}} | |||
| The fossils of multiple distinct Denisovan individuals from ] have been identified through their ] (aDNA): Denisova 2, 3, 4, 8, ], and 25. An mtDNA-based phylogenetic analysis of these individuals suggested that Denisova 2 is the oldest, followed by Denisova 8, while Denisova 3 and Denisova 4 were roughly contemporaneous.<ref name="Slon Viola Renaud Gansauge 2017"/> In 2024, scientists announced the sequence of Denisova 25, which was in a layer dated to 200ka.<ref name="h523">{{cite journal | title=The most ancient human genome yet has been sequenced—and it's a Denisovan's | last=Gibbons | first=Ann | date=2024-07-11 | url=https://www.science.org/content/article/most-ancient-human-genome-yet-has-been-sequenced-and-it-s-denisovan | doi=10.1126/science.zi9n4zp | journal=Science | access-date=13 July 2024 | archive-date=18 July 2024 | archive-url=https://web.archive.org/web/20240718134139/https://www.science.org/content/article/most-ancient-human-genome-yet-has-been-sequenced-and-it-s-denisovan | url-status=live }}</ref> During DNA sequencing, a low proportion of the Denisova 2, Denisova 4 and Denisova 8 genomes were found to have survived, but a high proportion of the Denisova 3 and Denisova 25 genomes were intact.<ref name="Slon Viola Renaud Gansauge 2017"/><ref name="PNAS-20151111">{{cite journal |doi=10.1073/pnas.1519905112 |pmid=26630009 |pmc=4697428 |title=Nuclear and mitochondrial DNA sequences from two Denisovan individuals |journal=] |volume=112 |issue=51 |pages=15696–700 |year=2015 |last1=Sawyer |first1=S. |last2=Renaud |first2=G. |last3=Viola |first3=B. |last4=Hublin |first4=J.-J. |last5=Gansauge |first5=M.-T. |last6=Shunkov |first6=M. V. |last7=Derevianko |first7=A. P. |last8=Prüfer |first8=K. |last9=Kelso |first9=J. |last10=Pääbo |first10=S. |author10-link=Svante Pääbo |display-authors=4 |bibcode=2015PNAS..11215696S |doi-access=free}}</ref><ref name="h523"/> The Denisova 3 sample was cut into two, and the initial DNA sequencing of one fragment was later independently confirmed by sequencing the mtDNA from the second.<ref name=Bennett /> | |||
| Denisova Cave contained the only known examples of Denisovans until 2019, when a research group led by ], ], and ] described a partial mandible discovered in 1980 by a ] in the ] on the ] in China. Known as the ], the fossil became part of the collection of ], where it remained unstudied until 2010.<ref name=Gibbon2019>{{cite journal|title=First fossil jaw of Denisovans finally puts a face on elusive human relatives|last=Gibbons|first=Anne|year=2019|journal=Science|doi=10.1126/science.aax8845|s2cid=188493848}}</ref> It was determined by ] analysis to contain ] that by sequence was found to have close affiliation to that of the Denisovans from Denisova Cave, while ] dating of the ] crust enshrouding the specimen indicated it was more than 160,000 years old.<ref name=ChenBaishiya>{{cite journal |first1=F. |last1=Chen |first2=F. |last2=Welker |first3=C.-C. |last3=Shen |display-authors=et al. |year=2019 |title=A late Middle Pleistocene Denisovan mandible from the Tibetan Plateau |journal=] |volume=569 |issue=7756 |pages=409–412 |doi=10.1038/s41586-019-1139-x |pmid=31043746 |author1-link=Fahu Chen |url=https://kar.kent.ac.uk/74280/1/Xiahe_Main.pdf |bibcode=2019Natur.569..409C |s2cid=141503768 |access-date=7 December 2019 |archive-date=13 December 2019 |archive-url= https://web.archive.org/web/20191213105210/https://kar.kent.ac.uk/74280/1/Xiahe_Main.pdf |url-status=live}}</ref> The identity of this population was later confirmed through study of ], which found Denisovan mtDNA in sediment layers ranging in date from 100,000 to 60,000 years before present, and perhaps more recent.<ref>{{cite journal|last=Shang |first=D. |display-authors=et al.|title=Denisovan DNA in Late Pleistocene sediments from Baishiya Karst Cave on the Tibetan Plateau |journal=] |volume=370 |number=6516 |pages=584–587 |year=2020 |doi=10.1126/science.abb6320 |pmid=33122381 |s2cid=225956074}}</ref> A 2024 reanalysis identified a partial Denisovan rib fragment dating to between 48,000 BP and 32,000 BP.<ref name=":2"/> | |||
| ==Mitochondrial DNA analysis== | |||
| The mtDNA from the finger bone differs from that of modern humans by 385 bases (]s) in the mtDNA strand out of approximately 16,500, whereas the difference between modern humans and ] is around 202 bases. In contrast, the difference between ]s and modern humans is approximately 1,462 mtDNA base pairs.<ref name="Pääbo et al." /> This suggested a divergence time around one million years ago. The mtDNA from a tooth bore a high similarity to that of the finger bone, indicating they belonged to the same population.<ref name="Reich et al."/> From a second tooth, an mtDNA sequence was recovered that showed an unexpectedly large number of genetic differences compared to that found in the other tooth and the finger, suggesting a high degree of mtDNA diversity. These two individuals from the same cave showed more diversity than seen among sampled Neanderthals from all of Eurasia, and were as different as modern-day humans from different continents.<ref name=Pennisi /> | |||
| In 2018, a team of Laotian, French, and American anthropologists, who had been excavating caves in the Laotian jungle of the ] since 2008, was directed by local children to the site Tam Ngu Hao 2 ("Cobra Cave") where they recovered a human tooth. The tooth (catalogue number TNH2-1) developmentally matches a 3.5 to 8.5 year old, and a lack of ] (a protein on the ]) suggests it belonged to a girl barring extreme degradation of the protein over a long period of time. Dental proteome analysis was inconclusive for this specimen, but the team found it anatomically comparable with the Xiahe mandible, and so tentatively categorized it as a Denisovan, although they could not rule out it being Neanderthal. The tooth probably dates to 164,000 to 131,000 years ago.<ref name=Demeter2022>{{cite journal|first1=F. |last1=Demeter |first2=C. |last2=Zanolli |first3=K. E. |last3=Westaway |display-authors=et al. |year=2022 |title=A Middle Pleistocene Denisovan molar from the Annamite Chain of northern Laos |journal=] |volume=13 |issue=2557 |page=2557 |doi=10.1038/s41467-022-29923-z |pmid=35581187 |pmc=9114389 |bibcode=2022NatCo..13.2557D }}</ref> | |||
| ==Nuclear genome analysis== | |||
| In the same second 2010 paper, the authors reported the isolation and sequencing of nuclear DNA from the Denisova finger bone. This specimen showed an unusual degree of DNA preservation and low level of contamination. They were able to achieve near-complete genomic sequencing, allowing a detailed comparison with Neanderthal and modern humans. From this analysis, they concluded, in spite of the apparent divergence of their mitochondrial sequence, the Denisova population along with Neanderthal shared a common branch from the lineage leading to modern African humans. The estimated average time of divergence between Denisovan and Neanderthal sequences is 640,000 years ago, and the time between both of these and the sequences of modern Africans is 804,000 years ago. They suggest the divergence of the Denisova mtDNA results either from the persistence of a lineage purged from the other branches of humanity through ] or else an ] from an older hominin lineage.<ref name="Reich et al."/> In 2013, the mtDNA sequence from the femur of a 400,000 year old '']'' from the ] in Spain was found to be most similar to that of Denisova.<ref name="NAT-20131204" /> | |||
| Some older findings may or may not belong to the Denisovan line, but Asia is not well mapped in regards to ]. Such findings include the ],<ref name=Callaway2010/> the ] hominin,<ref>{{cite journal|first1=H. |last1=Ao |first2=C.-R. |last2=Liu |first3=A. P. |last3=Roberts |year=2017 |title=An updated age for the Xujiayao hominin from the Nihewan Basin, North China: Implications for Middle Pleistocene human evolution in East Asia |journal=] |volume=106 |pages=54–65 |doi=10.1016/j.jhevol.2017.01.014 |pmid=28434540 |bibcode=2017AGUFMPP13B1080A |doi-access=free|hdl=1885/232536 |hdl-access=free }}</ref> ], the ] hominin, and the ].<ref name="Cooper"/> The Xiahe mandible shows morphological similarities to some later East Asian fossils such as ],<ref name=ChenBaishiya /><ref name=Warren2019>{{cite journal |first=M. |last=Warren |title=Biggest Denisovan fossil yet spills ancient human's secrets |journal=] |volume=569 |pages=16–17 |year=2019 |issue=7754 |doi=10.1038/d41586-019-01395-0 |pmid=31043736 |bibcode=2019Natur.569...16W |doi-access=free}}</ref> but also to Chinese ''H. erectus''.<ref name=Bennett>{{cite journal |first1=E. A. |last1=Bennett |first2=I. |last2=Crevecoeur |first3=B. |last3=Viola |display-authors=et al. |title= Morphology of the Denisovan phalanx closer to modern humans than to Neanderthals |journal=] |year=2019 |volume=5 |number=9 |page=eaaw3950 |doi=10.1126/sciadv.aaw3950 |pmid=31517046 |pmc=6726440 |bibcode=2019SciA....5.3950B}}</ref> In 2021, Chinese palaeoanthropologist Qiang Ji suggested his newly erected species, '']'', may represent the Denisovans based on the similarity between the type specimen's molar and that of the Xiahe mandible.<ref name= Ji2021>{{Cite journal |last1=Ji |first1=Qiang |last2=Wu |first2=Wensheng |last3=Ji |first3=Yannan |last4=Li |first4=Qiang |last5=Ni |first5=Xijun |display-authors=4 |date=2021-06-25 |title=Late Middle Pleistocene Harbin cranium represents a new Homo species |journal=The Innovation |volume=2 |issue=3 |page=100132 |language=English |doi=10.1016/j.xinn.2021.100132 |pmid=34557772 |pmc=8454552 |bibcode=2021Innov...200132J |issn=2666-6758 |doi-access=free}}</ref> In 2024, Bae and Wu suggested classifying the Xujiayao and Denisovan material as ''H. juluensis'', and the Dali Man and similar specimens as ''H. longi.<ref name=Bae2024/> | |||
| ==Interbreeding== | |||
| {{see also|Archaic human admixture with modern humans}} | |||
| ]A detailed comparison of the Denisovan, Neanderthal, and human genomes has revealed evidence for a complex web of interbreeding among the lineages. Through such interbreeding, 17% of the Denisova genome represents DNA from the local Neanderthal population, while evidence was also found of a contribution to the nuclear genome from an ancient hominin lineage yet to be identified,<ref name=Pennisi /> perhaps the source of the anomalously ancient mtDNA. | |||
| {| align="top" class="sortable wikitable" | |||
| Analysis of genomes of ] show that they mated with at least two groups of ]: ] (more similar to those found in the Caucasus than those from the Altai region)<ref name=Pennisi /> and Denisovans.<ref name="NYT-01302012" /><ref name="Reich et al."/><ref>{{cite journal |author=Green RE, Krause J, Briggs AW, et al. |title=A draft sequence of the Neandertal genome |journal=Science |volume=328 |issue=5979 |pages=710–22 |date=May 2010 |pmid=20448178 |doi=10.1126/science.1188021 |url=http://www.eva.mpg.de/neandertal/press/presskit-neandertal/pdf/Science_Green.pdf}}</ref> Approximately 4% of the ] of non-African ]s is shared with Neanderthals, suggesting interbreeding.<ref name="Reich et al."/> Tests comparing the Denisova hominin genome with those of six modern humans – a ] from South Africa, a ], a ], a ]n, a ]er and a ] – showed that between 4% and 6% of the ] of ] (represented by the Papua New Guinean and Bougainville Islander) derives from a Denisovan population. This DNA was possibly introduced during the early ] to Melanesia. These findings are in concordance with the results of other comparison tests which show a relative increase in allele sharing between the Denisovan and the Aboriginal Australian genome, compared to other Eurasians and African populations, however it has been observed that Papuans, the population of Papua New Guinea, have more allele sharing than Aboriginal Australians.<ref>Rasmussen et al 2011 An Aboriginal Australian genome reveals separate human dispersals into Asia. Science. 2011 Oct 7;334(6052):94-8. {{DOI|10.1126/science.1211177}}</ref> | |||
| |- | |||
| ! Name | |||
| ! Fossil elements | |||
| ! Age | |||
| ! Discovery | |||
| ! Place | |||
| ! Sex and age | |||
| ! Publication | |||
| ! Image | |||
| ! GenBank accession | |||
| |- | |||
| | Denisova 3<br />(also known as ''X Woman'')<ref name=pmid21179161/><ref name=Bennett /><ref name="Pääbo et al." /> || Distal ] of the ]|| 76.2–51.6 ka<ref name=Douka/> || align=center|2008 || Denisova cave (Russia) || 13.5-year-old adolescent female || align=center|2010 || ] || | |||
| |- | |||
| | Denisova 4<ref name=pmid21179161/><ref name=Callaway2010>{{cite journal |doi=10.1038/4681012a |pmid=21179140 |title=Fossil genome reveals ancestral link |journal=] |volume=468 |issue=7327 |pages=1012 |year=2010 |last1=Callaway |first1=Ewen |bibcode=2010Natur.468.1012C |doi-access=free}}</ref> || Permanent upper 2nd or 3rd molar || 84.1–55.2 ka<ref name=Douka/> || align=center|2000 || Denisova cave (Russia) || Adult male || align=center|2010 || ] | |||
| | | |||
| |- | |||
| | Denisova 8<ref name="PNAS-20151111"/> | |||
| || Permanent upper 3rd molar || 136.4–105.6 ka<ref name=Douka/> || align=center|2010 || Denisova cave (Russia) || Adult male || align=center|2015 || || | |||
| |- | |||
| | Denisova 2<ref name="Slon Viola Renaud Gansauge 2017">{{cite journal |doi=10.1126/sciadv.1700186 |pmid=28695206 |pmc=5501502 |title=A fourth Denisovan individual |journal=] |volume=3 |issue=7 |pages=e1700186 |year=2017 |author1-link=Viviane Slon |last1=Slon |first1=V. |last2=Viola |first2=B. |last3=Renaud |first3=G. |last4=Gansauge |first4=M.-T. |last5=Benazzi |first5=S. |last6=Sawyer |first6=S. |last7=Hublin |first7=J.-J. |last8=Shunkov |first8=M. V. |last9=Derevianko |first9=A. P. |last10=Kelso |first10=J. |last11=Prüfer |first11=K. |last12=Meyer |first12=M. |last13=Pääbo |first13=S. |author13-link=Svante Pääbo |bibcode=2017SciA....3E0186S |display-authors=4}}</ref> | |||
| || ] 2nd lower molar | |||
| || 194.4–122.7 ka<ref name=Douka/> || align=center|1984 || Denisova cave (Russia) || Adolescent female || align=center|2017 || || | |||
| |- | |||
| | ]<ref name=ChenBaishiya /> | |||
| | Partial mandible | |||
| | > 160 ka || align=center|1980 || ] (China) || || align=center|2019 ||] | |||
| | | |||
| |- | |||
| | ]<br />(also known as ''Denny'', Denisovan x Neanderthal hybrid)<br /><ref name="Brown 2016">{{cite journal|last1=Brown |first1=S. |last2=Higham |first2=T. |last3=Slon |first3=V. |last4=Pääbo |first4=S. |author4-link=Svante Pääbo |title=Identification of a new hominin bone from Denisova Cave, Siberia using collagen fingerprinting and mitochondrial DNA analysis |journal=] |year=2016 |volume=6 |doi=10.1038/srep23559 |page=23559 |pmid=27020421 |pmc=4810434 |bibcode=2016NatSR...623559B}}</ref> | |||
| | Arm or leg bone fragment | |||
| | 118.1–79.3 ka<ref name=Douka/> || align=center|2012 || Denisova cave (Russia) || 13 year old adolescent female || align=center|2016 ||] | |||
| | | |||
| |- | |||
| | Denisova 13<ref name=Viola2019>{{cite journal|first1=B. T. |last1=Viola |first2=P. |last2=Gunz |first3=S. |last3=Neubauer |year=2019 |title=A parietal fragment from Denisova cave |journal=88th Annual Meeting of the American Association of Physical Anthropologists |url=https://meeting.physanth.org/program/2019/session09/viola-2019-a-parietal-fragment-from-denisova-cave.html |access-date=18 January 2020 |archive-date=26 September 2019 |archive-url=https://web.archive.org/web/20190926091143/https://meeting.physanth.org/program/2019/session09/viola-2019-a-parietal-fragment-from-denisova-cave.html |url-status=live}}</ref> | |||
| | ] fragment | |||
| | Found in layer 22<ref name=Viola2019/> which dates to ~285±39 ka<ref name=Jacobs2019/> || align=center|2019 || Denisova cave (Russia) || ||align=center|pending | |||
| | | |||
| | | |||
| |- | |||
| | TNH2-1<ref name=Demeter2022/> | |||
| | Permanent lower left 1st or 2nd molar | |||
| | 164–131 ka || align=center|2018 || Tam Ngu Hao 2 cave (Laos) || 3.5 to 8.5 year old female || align=center|2022 ||] | |||
| | | |||
| |- | |||
| |BSY-19-B896-1 (Xiahe 2) | |||
| | Distal rib fragment | |||
| | 48–32 ka || align=center|1980 || Baishiya Cave (China) || Unknown || align=center|2024 || ] | |||
| |- | |||
| | Denisova 25<ref name="h523"/> | |||
| | Molar | |||
| | 200 ka || align=center|2024|| Denisova cave (Russia) || Male|| align=center|pending|| | |||
| |} | |||
| === Evolution === | |||
| Melanesians may not be the only modern-day descendants of Denisovans. David Reich of ], in collaboration with Mark Stoneking of the Planck Institute team, found genetic evidence that Denisovan ancestry is shared by Melanesians, ], and smaller scattered groups of people in Southeast Asia, such as the ], a ] people in the ]. However, not all ] were found to possess Denisovan genes; ] ] and Malaysian ], for example, were found to have no significant Denisovan inheritance. These data place the interbreeding event in mainland Southeast Asia, and suggest that Denisovans once ranged widely over eastern Asia.<ref name="Callaway."/><ref name="Reich">{{Citation|author=Reich ''et al.''|journal= | |||
| ]s, '']'' and '']'']] | |||
| The American Journal of Human Genetics|volume=89 |pages=516–28 |year=2011| title=Denisova Admixture and the First Modern Human Dispersals into Southeast Asia and Oceania|url=http://www.sciencedirect.com/science/article/pii/S0002929711003958 | doi=10.1016/j.ajhg.2011.09.005 | pmid=21944045 | pmc=3188841}}</ref><ref name="Choi.">{{Citation |url=http://www.livescience.com/16171-denisovans-humans-widespread-sex-asia.html |title=Now-Extinct Relative Had Sex with Humans Far and Wide |publisher=] |date=22 September 2011 |first=Charles |last=Choi}}</ref> Based on the modern distribution of Denisova DNA, Denisovans may have crossed the ], with ] serving as their last ].<ref name="Cooper">{{Citation|author=Cooper A. and Stringer C.B.|journal= | |||
| ] ] (mtDNA), preserved by the cool climate of the cave (average temperature is at freezing point), was extracted from Denisova 3 by a team of scientists led by ] and ] from the ] in ], Germany. Denisova 3's mtDNA differs from that of modern humans by 385 bases (]s) out of approximately 16,500, whereas the difference between modern humans and ] is around 202 bases. In comparison, the difference between ]s and modern humans is approximately 1,462 mtDNA base pairs. This suggested that Denisovan mtDNA diverged from that of modern humans and Neanderthals about 1,313,500–779,300 years ago; whereas modern human and Neanderthal mtDNA diverged 618,000–321,200 years ago. Krause and colleagues then concluded that Denisovans were the descendants of an earlier migration of ''H. erectus'' out of Africa, completely distinct from modern humans and Neanderthals.<ref name="Pääbo et al." /> | |||
| Science|volume=342 |issue=6156|pages=321–3|year=2013| title=Did the Denisovans Cross the Wallace Line|url=http://www.sciencemag.org/content/342/6156/321.summary | doi=10.1126/science.1244869 }}</ref><ref>http://www.abc.net.au/science/articles/2013/10/18/3869503.htm</ref> A paper by Kay Prüfer in 2013 said that mainland Asians and Native Americans had around 0.2% Denisovan ancestry.<ref name="NAT-2013">{{cite journal |author=Prüfer, Kay et al. |title=The complete genome sequence of a Neanderthal from the Altai Mountains |url=http://www.nature.com/nature/journal/v505/n7481/full/nature12886.html |journal=] |volume=505 |pages=43–49 |year=2013 |accessdate=1 March 2014 |doi=10.1038/nature12886}}</ref> | |||
| However, according to the ] (nDNA) of Denisova 3—which had an unusual degree of DNA preservation with only low-level contamination—Denisovans and Neanderthals were more closely related to each other than they were to modern humans. Using the percent distance from ], Denisovans/Neanderthals split from modern humans about 804,000 years ago, and from each other 640,000 years ago.<ref name=pmid21179161>{{cite journal |doi=10.1038/nature09710 |pmid=21179161 |pmc=4306417 |title=Genetic history of an archaic hominin group from Denisova Cave in Siberia |journal=] |volume=468 |issue=7327 |pages=1053–60 |year=2010 |last1=Reich |first1=D. |author1-link=David Reich (geneticist) |last2=Green |first2=R. E. |last3=Kircher |first3=M. |display-authors=et al. |url=http://repositori.upf.edu/bitstream/10230/25596/1/Marques_nat_gen.pdf |bibcode=2010Natur.468.1053R |hdl=10230/25596 |access-date=29 July 2018 |archive-date=17 May 2020 |archive-url=https://web.archive.org/web/20200517024908/https://repositori.upf.edu/bitstream/handle/10230/25596/Marques_nat_gen.pdf;jsessionid=35CCDAE27CDE1B5CACE38275BC976DFA?sequence=1 |url-status=live}}</ref> Using a mutation rate of {{val|1|e=-9}} or {{val|0.5|e=-9}} per ] (bp) per year, the Neanderthal/Denisovan split occurred around either 236–190,000 or 473–381,000 years ago respectively.<ref name=":0" /> Using {{val|1.1|e=-8}} per generation with a new generation every 29 years, the time is 744,000 years ago. Using {{val|5|e=-10}} ] site per year, it is 616,000 years ago. Using the latter dates, the split had likely already occurred by the time hominins spread out across Europe.<ref name=rogers2017>{{cite journal|first1=A. R. |last1=Rogers |first2=R. J. |last2=Bohlender |first3=C. D. |last3=Huff |title=Early history of Neanderthals and Denisovans |journal=] |volume=114 |issue=37 |year=2017 |pages=9859–9863 |doi=10.1073/pnas.1706426114 |pmid=28784789 |pmc=5604018 |bibcode=2017PNAS..114.9859R |doi-access=free}}</ref> '']'' is typically considered to have been the direct ancestor of Denisovans and Neanderthals, and sometimes also modern humans.<ref>{{cite journal|first=K. K. |last=Ho |year=2016 |title=Hominin interbreeding and the evolution of human variation |journal=Journal of Biological Research-Thessaloniki |volume=23 |page=17 |doi=10.1186/s40709-016-0054-7 |pmc=4947341 |pmid=27429943 |doi-access=free }}</ref> Due to the strong divergence in dental anatomy, they may have split before characteristic Neanderthal dentition evolved about 300,000 years ago.<ref name=pmid21179161/> | |||
| The immune system's ] alleles have drawn particular attention in the attempt to identify genes that may derive from archaic human populations. Although not present in the sequenced Denisova genome, the distribution pattern and divergence of HLA-B*73 from other HLA alleles has led to the suggestion that it ] from Denisovans into humans in west Asia. Indeed, half of the HLA alleles of modern Eurasians represent archaic HLA haplotypes, and have been inferred to be of Denisovan or Neanderthal origin.<ref name="10.1126/science.1209202">{{cite journal | author=Laurent Abi-Rached, et al. | title= The Shaping of Modern Human Immune Systems by Multiregional Admixture with Archaic Humans | journal=Science | volume=334 | issue=6052 | pages= | date=25 August 2011 |doi=10.1126/science.1209202 |archiveurl=http://digitalcommons.unl.edu/cgi/viewcontent.cgi?article=1122&context=publichealthresources|archivedate=Aug 2011|pmc=3677943| url=http://www.sciencemag.org/content/early/2011/08/19/science.1209202 | pmid=21868630 | |||
| |bibcode = |laysummary=http://www.bbc.co.uk/news/science-environment-14673047 }}</ref> The apparent over-representation of these alleles suggests a positive selective pressure for their retention in the human population. A higher quality Denisovan genome published in 2012 reveals variants of genes in humans that are associated with dark skin, brown hair and brown eyes - consistent with features found with Melanesians today.<ref>Marshall, Michael (2014), op cit, p.38</ref> | |||
| The more divergent Denisovan mtDNA has been interpreted as evidence of admixture between Denisovans and an unknown archaic human population,<ref>{{cite journal|last=Malyarchuk|first=B. A.|date =2011|title=Adaptive evolution of the ''Homo'' mitochondrial genome|journal= Molecular Biology|volume= 45|issue=5|pages =845–850|doi=10.1134/S0026893311050104|pmid=22393781 |s2cid=43284294}}</ref> possibly a ] '']'' or ''H. erectus''-like population about 53,000 years ago.<ref name=":0">{{Cite journal |last1=Lao |first1=O. |last2=Bertranpetit |first2=J. |last3=Mondal |first3=M. |year=2019 |title=Approximate Bayesian computation with deep learning supports a third archaic introgression in Asia and Oceania |journal=] |language=en |volume=10 |issue=1 |pages=246 |doi=10.1038/s41467-018-08089-7 |issn=2041-1723 |pmc=6335398 |pmid=30651539 |bibcode=2019NatCo..10..246M}}</ref> Alternatively, divergent mtDNA could have also resulted from the persistence of an ancient mtDNA lineage which only went extinct in modern humans and Neanderthals through ].<ref name=pmid21179161/> Modern humans contributed mtDNA to the Neanderthal lineage, but not to the Denisovan mitochondrial genomes yet sequenced.<ref>{{Cite journal|last1=Pääbo |first1=S. |author1-link=Svante Pääbo |last2=Kelso |first2=J. |last3=Reich |first3=D. |author3-link=David Reich (geneticist) |last4= Slatkin |first4=M. |last5=Viola |first5=B. |last6=Derevianko |first6=A. P. |last7=Shunkov |first7=M. V. |last8=Doronichev |first8=V. B. |last9=Golovanova |first9= L. V. |display-authors=4 |year= 2014 |title=The complete genome sequence of a Neanderthal from the Altai Mountains |journal=] |language=en |volume=505 |issue=7481 |pages=43–49 |doi=10.1038/nature12886 |issn=1476-4687 |pmc=4031459 |pmid=24352235 |bibcode=2014Natur.505...43P}}</ref><ref>{{Cite journal |last1=Kuhlwilm |first1=M. |last2=Gronau |first2=I. |last3=Hubisz |first3=M. J. |last4=de Filippo |first4=C. |last5=Prado-Martinez |first5=J. |last6=Kircher |first6=M. |last7=Fu |first7=Q. |last8=Burbano |first8=H. A. |last9=Lalueza-Fox |first9=C. |display-authors=4 |year=2016 |title=Ancient gene flow from early modern humans into Eastern Neanderthals |journal=] |volume=530 |issue=7591 |pages=429–433 |doi=10.1038/nature16544 |issn=1476-4687 |pmc=4933530 |pmid=26886800 |bibcode=2016Natur.530..429K}}</ref><ref>{{Cite journal| last1= Posth| first1= C. |last2=Wißing |first2=C. |last3=Kitagawa |first3=K. |last4=Pagani |first4=L. |last5=van Holstein |first5=L. |last6=Racimo |first6=F. |last7=Wehrberger |first7=K.|last8=Conard |first8=N. J. |last9=Kind |first9=C. J. |display-authors=4 |year=2017 |title=Deeply divergent archaic mitochondrial genome provides lower time boundary for African gene flow into Neanderthals |journal=] |volume=8 |pages=16046 |doi=10.1038/ncomms16046 |issn=2041-1723 |pmc=5500885 |pmid=28675384 |bibcode= 2017NatCo...816046P}}</ref><ref>{{Cite journal|last1=Bertranpetit |first1=J. |last2=Majumder |first2=P. P. |last3=Li |first3=Q. |last4=Laayouni |first4=H. |last5=Comas |first5=D. |last6=Netea |first6=M. G. |last7=Pybus |first7=M. |last8=Dall'Olio |first8=G. M. |last9=Xu |first9=T. |display-authors=4 |year=2016 |title=Genomic analysis of Andamanese provides insights into ancient human migration into Asia and adaptation |journal=] |volume=48 |issue=9 |pages=1066–1070 |doi=10.1038/ng.3621 |pmid=27455350 |issn=1546-1718 |hdl=10230/34401 |s2cid=205352099 |hdl-access=free}}</ref> The mtDNA sequence from the femur of a 400,000-year-old ''H. heidelbergensis'' from the ] in Spain was found to be related to those of Neanderthals and Denisovans, but closer to Denisovans,<ref name="NAT-20131204">{{cite journal |doi=10.1038/504016a |pmid=24305130 |title=Hominin DNA baffles experts |journal=] |volume=504 |issue=7478 |pages=16–17 |year=2013 |last1=Callaway |first1= E. |bibcode= 2013Natur.504...16C |doi-access=free }}</ref><ref>{{cite book|first=I. |last=Tattersall |author-link=Ian Tattersall |title=The Strange Case of the Rickety Cossack and other Cautionary Tales from Human Evolution |page=200 |publisher=] |year=2015 |isbn=978-1-137-27889-0}}</ref> and the authors posited that this mtDNA represents an archaic sequence which was subsequently lost in Neanderthals due to replacement by a modern-human-related sequence.<ref name="Meyer2016">{{cite journal |first1=M. |last1=Meyer |first2=J.-L. |last2=Arsuaga |display-authors=et al. |title=Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins |journal=] |volume=531 |issue=7595 |pages=504–507 |year=2016 |doi=10.1038/nature17405 |pmid=26976447 |bibcode=2016Natur.531..504M |s2cid=4467094}}</ref> | |||
| It has been suggested,<ref>{{cite web|last=Barras|first=Colin|title=Chinese fossils unlike any known species|url=http://www.newscientist.com/article/dn21586|publisher=New Scientist (Reed Business Information)|accessdate=19 December 2013}}</ref> in the absence of genomic evidence (as of 2013), that the ] of China were the result of interbreeding between Homo sapiens and Denisovans within a few thousands years of the end of the ]. | |||
| == Demographics == | |||
| There is evidence of a minimum 0.5% Neanderthal gene flow into the Denisovans.<ref name=prualtai>{{cite journal| last=Prüfer|first=K.|author2=Racimo, Fe. |author3=Patterson, N. |author4=Jay, F. |author5=Sankararaman, S. |author6=Sawyer, S. |author7= et al. | title=The complete genome sequence of a Neanderthal from the Altai Mountains| journal=Nature| year=2013| volume=505| issue=7481| pages=43–49| doi=10.1038/nature12886}}</ref> The Denisovan genome shared more derived alleles with the Altai Neanderthal genome from Siberia than with the Vindija Neanderthal genome from Croatia and the Mezmaiskaya Neanderthal genome from the Caucasus, suggesting that the gene flow came from a population that was more closely related to the Altai Neanderthal.<ref name=prualtai/> | |||
| {{See also|Archaic humans in Southeast Asia}} | |||
| ].<ref name="Cooper"/>]] | |||
| Denisovans are known to have lived in Siberia, Tibet, and Laos.<ref name=Demeter2022/> The Xiahe mandible is the earliest recorded human presence on the Tibetan Plateau.<ref name=ChenBaishiya/> Though their remains have been identified in only these three locations, traces of Denisovan DNA in modern humans suggest they ranged across ],<ref name= "Callaway2011">{{cite journal |doi=10.1038/news.2011.551 |title=First Aboriginal genome sequenced |journal=] |year=2011 |last1=Callaway |first1=E.}}</ref><ref name="Reich"/> and potentially western Eurasia.<ref name=Skov2020/> In 2019, ] Guy Jacobs and colleagues identified three distinct populations of Denisovans responsible for the introgression into modern populations now native to, respectively: Siberia and East Asia; New Guinea and nearby islands; and ] and, to a lesser extent, across Asia. Using ], the Denisova Cave Denisovans split from the second population about 283,000 years ago; and from the third population about 363,000 years ago. This indicates that there was considerable reproductive isolation between Denisovan populations.<ref name=JacobsG2019>{{Cite journal |last1=Jacobs |first1=G. S. |last2=Hudjashov |first2=G. |last3=Saag |first3=L. |last4=Kusuma |first4=P. |last5=Darusallam |first5=C. C. |last6=Lawson |first6=D. J. |last7=Mondal |first7=M. |last8=Pagani |first8=L. |last9=Ricaut |first9= F.-X. |last10=Stoneking |first10=M. |last11=Metspalu |first11=M. |display-authors=4 |year=2019 |title=Multiple Deeply Divergent Denisovan Ancestries in Papuans |journal=] |volume=177 |issue=4 |pages=1010–1021.e32 |doi=10.1016/j.cell.2019.02.035 |pmid=30981557 |issn=0092-8674 |doi-access=free|hdl=1983/7df38af7-d075-4444-9111-b859650f6d38 |hdl-access=free }}</ref> In a 2024 study, scientist Danat Yermakovich, of the ], discovered that people living at different elevations in Papua New Guinea have differences in Denisovan DNA; with people living in the highlands having variants for early brain development and those living in the lowlands having variants for the immune system.<ref>{{cite web | url=https://www.newscientist.com/article/2438941-denisovan-dna-may-help-modern-humans-adapt-to-different-environments/ | title=Denisovan DNA may help modern humans adapt to different environments | access-date=11 August 2024 | archive-date=11 August 2024 | archive-url=https://web.archive.org/web/20240811223304/https://www.newscientist.com/article/2438941-denisovan-dna-may-help-modern-humans-adapt-to-different-environments/ | url-status=live }}</ref> | |||
| Based on the high percentages of Denisovan DNA in modern Papuans and Australians, Denisovans may have crossed the ] into these regions (with little back-migration west), the second known human species to do so,<ref name="Cooper">{{cite journal |doi=10.1126/science.1244869 |pmid=24136958 |title=Did the Denisovans Cross Wallace's Line? |journal=] |volume=342 |issue=6156 |pages=321–23 |year=2013 |last1=Cooper |first1=A. |last2=Stringer |first2=C. B. |author2-link=Chris Stringer |bibcode= 2013Sci...342..321C |s2cid=206551893}}</ref> along with earlier '']''. By this logic, they may have also entered the Philippines, living alongside '']'' which, if this is the case, may represent the same or a closely related species.<ref name=Larena2021/> These Denisovans may have needed to cross large bodies of water.<ref name= JacobsG2019/> Alternately, high Denisovan DNA admixture in modern Papuan populations may simply represent higher mixing among the original ancestors of Papuans prior to crossing the Wallace line. Icelanders also have an anomalously high Denisovan heritage, which could have stemmed from a Denisovan population far west of the Altai Mountains. Genetic data suggests Neanderthals were frequently making long crossings between Europe and the Altai Mountains especially towards the date of their extinction.<ref name= Skov2020/> | |||
| It has also been observed that the Denisovan genome comprises a component derived from an unknown hominin that diverged long before the modern human/Neanderthal/Denisovan separated, suggesting a possible gene flow from said unknown hominin to Denisovans or a population sub-structure.<ref name=prualtai/> | |||
| Using ] analysis on ] lengths, Jacobs calculated ] into modern humans occurred about 29,900 years ago with the Denisovan population ancestral to New Guineans; and 45,700 years ago with the population ancestral to both New Guineans and Oceanians. Such a late date for the New Guinean group could indicate Denisovan survival as late as 14,500 years ago, which would make them the latest surviving archaic human species. A third wave appears to have introgressed into East Asia, but there is not enough DNA evidence to pinpoint a solid timeframe.<ref name=JacobsG2019/> | |||
| Tibetans have a version of the ] gene which helps them to adapt to the low oxygen levels at high altitude, and according to a study published in ] in July 2014, this version of the gene is found in the Denisovan genome, suggesting that they acquired the adaptation by interbreeding between their ancestors and the Denisovans.<ref>{{cite journal|journal=Nature|doi=10.1038/nature13408|url=http://www.nature.com/nature/journal/vaop/ncurrent/full/nature13408.html|title=Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA|authors=Huerta-Sanchez, Emilia et al|date=2 July 2014}}; {{cite journal|journal=New Scientist|first=Catherine|last=Brahic|title=Extinct humans primed Tibetans for the high life|date=5 July 2014}}</ref> | |||
| The mtDNA from Denisova 4 bore a high similarity to that of Denisova 3, indicating that they belonged to the same population.<ref name=pmid21179161/> The ] among the Denisovans from Denisova Cave is on the lower range of what is seen in modern humans, and is comparable to that of Neanderthals. However, it is possible that the inhabitants of Denisova Cave were more or less reproductively isolated from other Denisovans, and that, across their entire range, Denisovan genetic diversity may have been much higher.<ref name="Slon Viola Renaud Gansauge 2017"/> | |||
| ==References== | |||
| Denisova Cave, over time of habitation, continually swung from a fairly warm and moderately humid pine and birch forest to tundra or forest-tundra landscape.<ref name=Jacobs2019/> Conversely, Baishiya Karst Cave is situated at a high elevation, an area characterized by low temperature, low oxygen, and poor resource availability. Colonization of high-altitude regions, due to such harsh conditions, was previously assumed to have only been accomplished by modern humans.<ref name=ChenBaishiya /> Denisovans seem to have also inhabited the jungles of Southeast Asia.<ref name="Reich"/> The Tam Ngu Hao 2 site might have been a closed forest environment.<ref name=Demeter2022/> | |||
| == Anatomy == | |||
| Little is known of the precise anatomical features of the Denisovans since the only physical remains discovered so far are a finger bone, four teeth, ] fragments, a partial jawbone,<ref name=Gibbon2019 /><ref name=Demeter2022/> a ] skull fragment,<ref name=Viola2019/> and a rib bone.<ref name=":2" /> The finger bone is within the modern human range of variation for women,<ref name=Bennett /> which is in contrast to the large, robust molars which are more similar to those of Middle to Late Pleistocene archaic humans. The third molar is outside the range of any ''Homo'' species except '']'' and '']'', and is more like those of ]s. The second molar is larger than those of modern humans and Neanderthals, and is more similar to those of '']'' and ''H. habilis''.<ref name=pmid21179161/> Like Neanderthals, the mandible had a gap behind the molars, and the front teeth were flattened; but Denisovans lacked a high mandibular body, and the ] at the midline of the jaw was more receding.<ref name=ChenBaishiya /><ref name=Warren2019/> The parietal is reminiscent of that of ''H. erectus''.<ref name=Callaway2019>{{cite journal|first=E. |last=Callaway |year=2019 |title=Siberia's ancient ghost clan starts to surrender its secrets |journal=] |volume=566 |issue=7745 |pages=444–446 |doi=10.1038/d41586-019-00672-2 |pmid=30814723 |bibcode=2019Natur.566..444C |doi-access=free}}</ref> | |||
| A facial reconstruction has been generated by comparing ] at individual genetic loci associated with facial structure.<ref name= 'Gokhman2014'>{{cite journal |last1=Gokhman |first1=D. |last2=Lavi |first2=E. |last3=Prüfer |first3=K. |last4=Fraga |first4=M. F. |last5=Riancho |first5=J. A. |last6=Kelso |first6= J |last7=Pääbo |first7=S. |author7-link=Svante Pääbo |last8=Meshorer |first8=E. |last9=Carmel |first9=L. |display-authors=4 |title=Reconstructing the DNA methylation maps of the Neandertal and the Denisovan |journal=] |volume=344 |issue=6183 |pages=523–27 |year=2014 |doi=10.1126/science.1250368 |pmid=24786081 |bibcode= 2014Sci...344..523G |s2cid=28665590|doi-access=free }}</ref> This analysis suggested that Denisovans, much like Neanderthals, had a long, broad, and projecting face; large nose; sloping forehead; protruding jaw; elongated and flattened skull; and wide chest and hips. The Denisovan ] was longer than that of Neanderthals and anatomically modern humans.<ref name="CELL-20190919">{{cite journal |last1=Gokhman |first1=D. |first2=N. |last2=Mishol |first3=M. |last3=de Manuel |first4=T. |last4=Marques-Bonet |first5=Y. |last5=Rak |first6=L. |last6=Carmel |display-authors=4 |title=Reconstructing Denisovan Anatomy Using DNA Methylation Maps |year=2019 |journal=] |volume=179 |issue=1 |pages=180–192 |doi=10.1016/j.cell.2019.08.035 |pmid=31539495 |s2cid=202676502 |doi-access=free}}</ref> | |||
| Middle-to-Late Pleistocene East Asian archaic human skullcaps typically share features with Neanderthals. The skullcaps from Xuchang feature prominent brow ridges like Neanderthals, though the nuchal and angular tori near the base of the skull are either reduced or absent, and the back of the skull is rounded off like in ]s. Xuchang 1 had a large brain volume of approximately 1800 cc, on the high end for Neanderthals and early modern humans, and well beyond the present-day human average.<ref>{{cite journal|first1=Z.-Y. |last1=Li |first2= X.-J. |last2=Wu |first3=L.-.P |last3=Zhou |display-authors=et al. |year=2017 |title=Late Pleistocene archaic human crania from Xuchang, China |journal=] |volume=355 |issue=6328 |pages=969–972 |doi=10.1126/science.aal2482 |pmid=28254945 |bibcode=2017Sci...355..969L |s2cid=206654741}}</ref> | |||
| The Denisovan genome from Denisova Cave has variants of genes which, in modern humans, are associated with dark skin, brown hair, and brown eyes.<ref name=Meyer2012>{{cite journal|first1=M. |last1=Meyer |first2=M. |last2=Kircher |first3=M.-T. |last3=Gansauge |display-authors=et al. |year=2012 |title=A High-Coverage Genome Sequence from an Archaic Denisovan Individual |journal=] |volume=338 |issue=6104 |pages=222–226 |doi=10.1126/science.1224344 |pmid=22936568 |pmc=3617501 |bibcode=2012Sci...338..222M}}</ref> The Denisovan genome also contains a variant region around the ] gene that in ] assists with adaptation to low oxygen levels at high elevation,<ref name=Huerta2014>{{cite journal |doi=10.1038/nature13408 |pmid=25043035 |pmc=4134395 |title=Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA |journal=] |volume=512 |issue=7513 |pages=194–97 |year=2014 |last1=Huerta-Sánchez |first1=E.|last2= Jin |first2=X. |display-authors=et al. |bibcode=2014Natur.512..194H}}</ref><ref name=ChenBaishiya /> and in a region containing the ] and ] loci which affect body-fat distribution in the ].<ref>{{cite journal|first1=Fernando |last1=Racimo |first2=David |last2=Gokhman |first3=Matteo |last3=Fumagalli |first4=Amy |last4=Ko |display-authors=4 |first5=Torben |last5=Hansen |first6=Ida |last6=Moltke |first7=Anders |last7=Albrechtsen |first8=Liran |last8=Carmel |author8-link=Liran Carmel |first9=Emilia |last9=Huerta-Sánchez |first10=Rasmus |last10=Nielsen |author10-link=Rasmus Nielsen (biologist) |title=Archaic Adaptive Introgression in TBX15/WARS2 |journal=] |volume=34 |number=3 |year=2017 |pages=509–524 |doi=10.1093/molbev/msw283 |pmid=28007980 |pmc=5430617}}</ref> In Papuans, introgressed Neanderthal alleles are highest in frequency in genes expressed in the brain, whereas Denisovan alleles have highest frequency in genes expressed in bones and other tissue.<ref>{{cite journal |doi=10.1142/S0219720018400115 |pmid=29739306 |title=Neanderthal and Denisovan ancestry in Papuans: A functional study |journal=] |volume=16 |issue=2 |pages=1840011 |year=2018 |last1=Akkuratov |first1=Evgeny E. |last2=Gelfand |first2=Mikhail S. |last3=Khrameeva |first3=Ekaterina E.}}</ref> | |||
| == Culture == | |||
| === Denisova Cave === | |||
| {{Multiple image|direction=vertical|width=250|image1=Denisova Cave pendants notched bones.jpg|image2=Denisova Cave lithic and osseous artifacts grey.jpg|footer=Some ornaments (above) and animal bones and stone tools (below) found in Denisova Cave. Note, ornaments may have been crafted by modern humans}} | |||
| Early ] stone tools from Denisova Cave included ], ]s, ]s, and notched tools, deposited about 287±41 thousand years ago in the Main Chamber of the cave; and about 269±97 thousand years ago in the South Chamber; up to 170±19 thousand and 187±14 thousand years ago in the Main and East Chambers, respectively.<ref name=Jacobs2019>{{Cite journal|last1=Jacobs |first1=Zenobia |last2=Li |first2=Bo |last3=Shunkov |first3=Michael V. |last4=Kozlikin |first4=Maxim B. |last5=Bolikhovskaya |first5=Nataliya S. |last6=Agadjanian |first6=Alexander K. |last7=Uliyanov |first7=Vladimir A. |last8=Vasiliev |first8=Sergei K. |last9=O’Gorman |first9=Kieran |last10=Derevianko |first10=Anatoly P. |last11=Roberts |first11=Richard G. |display-authors=4 |date=January 2019 |title=Timing of archaic hominin occupation of Denisova Cave in southern Siberia |journal=] |volume=565 |issue=7741 |pages=594–599 |doi=10.1038/s41586-018-0843-2 |pmid=30700870 |issn=1476-4687 |bibcode=2019Natur.565..594J |s2cid=59525956 |url=https://ro.uow.edu.au/cgi/viewcontent.cgi?article=1558&context=smhpapers1|access-date=29 May 2020 |archive-date=7 May 2020 |archive-url=https://web.archive.org/web/20200507004154/https://ro.uow.edu.au/cgi/viewcontent.cgi?article=1558&context=smhpapers1 |url-status=live}}</ref> | |||
| Middle Paleolithic assemblages were dominated by flat, discoidal, and Levallois cores, and there were some isolated sub-prismatic cores. There were predominantly side scrapers (a scraper with only the sides used to scrape), but also notched-denticulate tools, end-scrapers (a scraper with only the ends used to scrape), ], ]-like tools, and truncated flakes. These dated to 156±15 thousand years ago in the Main Chamber, 58±6 thousand years ago in the East Chamber, and 136±26–47±8 thousand years ago in the South Chamber.<ref name=Jacobs2019/> | |||
| Early ] artefacts date to 44±5 thousand years ago in the Main Chamber, 63±6 thousand years ago in the East Chamber, and 47±8 thousand years ago in the South Chamber, though some layers of the East Chamber seem to have been disturbed. There was ] production and Levallois production, but scrapers were again predominant. A well-developed, Upper Paleolithic stone bladelet technology distinct from the previous scrapers began accumulating in the Main Chamber around 36±4 thousand years ago.<ref name=Jacobs2019/> | |||
| In the Upper Paleolithic layers, there were also several ]s and ornaments: a ] ring, an ivory ring, an ivory pendant, a ] tooth pendant, an ] tooth pendant, a ] bracelet, and a bone needle. However, Denisovans are only confirmed to have inhabited the cave until 55 ka; the dating of Upper Paleolithic artefacts overlaps with modern human migration into Siberia (though there are no occurrences in the Altai region); and the DNA of the only specimen in the cave dating to the time interval (Denisova 14) is too degraded to confirm species identity, so the attribution of these artefacts is unclear.<ref>{{cite journal |first=R. |last=Dennell |year=2019 |title=Dating of hominin discoveries at Denisova |journal=] |volume=565 |issue=7741 |pages=571–572 |doi=10.1038/d41586-019-00264-0 |pmid=30700881 |bibcode=2019Natur.565..571D |doi-access=free}}</ref><ref name=Jacobs2019/> | |||
| === Tibet === | |||
| ]s from Baishiya Karst Cave: a) ], b) ], c) ], d) horse, e) ]]] | |||
| The inhabitants of Baishiya Karst Cave seem to have been extensively processing ], cows, deer, horses, and ]. They were also butchering large carnivores (], dog, and big cat), ]s, ], and ]s. They may have also used these animals' ]s to make bone tools, and additionally there are stone artefacts in each layer excavated.<ref name=":2">{{Cite journal |last1=Xia |first1=Huan |last2=Zhang |first2=Dongju |last3=Wang |first3=Jian |last4=Fagernäs |first4=Zandra |last5=Li |first5=Ting |last6=Li |first6=Yuanxin |last7=Yao |first7=Juanting |last8=Lin |first8=Dongpeng |last9=Troché |first9=Gaudry |last10=Smith |first10=Geoff M. |last11=Chen |first11=Xiaoshan |last12=Cheng |first12=Ting |last13=Shen |first13=Xuke |last14=Han |first14=Yuanyuan |last15=Olsen |first15=Jesper V. |date=3 July 2024 |title=Middle and Late Pleistocene Denisovan subsistence at Baishiya Karst Cave |journal=Nature |volume=632 |issue=8023 |language=en |pages=108–113 |doi=10.1038/s41586-024-07612-9 |issn=1476-4687|doi-access=free |pmid=38961285 |pmc=11291277 }}</ref> | |||
| In 1998, five child hand- and footprint impressions were discovered in a ] ] near the Quesang ]s in Tibet; in 2021, they were dated to 226 to 169 thousand years ago using uranium decay dating. This is the oldest evidence of human occupation of the Tibetan Plateau, and since the Xiahe mandible is the oldest human fossil from the region (though younger than the Quesang impressions), these may have been made by Denisovan children. The impressions were printed onto a small panel of space, and there is little overlap between all the prints, so they seem to have been taking care to make new imprints in unused space. If considered art, they are the oldest known examples of ]. Similar hand stencils and impressions do not appear again in the archeological record until roughly 40,000 years ago.<ref name=Zhang2021>{{cite journal |first1=D. D. |last1=Zhang |first2=M. R. |last2=Bennett |first3=H. |last3=Cheng |first4=L. |last4=Wang |display-authors=et al. |year=2021 |title=Earliest parietal art: Hominin hand and foot traces from the middle Pleistocene of Tibet |journal=] |volume=66 |issue=24 |pages=2506–2515 |doi=10.1016/j.scib.2021.09.001 |pmid=36654210 |bibcode=2021SciBu..66.2506Z |issn=2095-9273 |s2cid=239102132|doi-access=free }}</ref> | |||
| The footprints comprise four right impressions and one left superimposed on one of the rights. They were probably left by two individuals. The tracks of the individual who superimposed their left onto their right may have scrunched up their toes and wiggled them in the mud, or dug their finger into the toe prints. The footprints average {{cvt|192.3|mm}} long, which roughly equates to a 7 or 8 year old child by modern human growth rates. There are two sets of handprints (from a left and right hand), which may have been created by an older child unless one of the former two individuals had long fingers. The handprints average {{cvt|161.1|mm}}, which roughly equates with a 12 year old modern human child, and the middle finger length agrees with a 17 year old modern human. One of the handprints shows an impression of the forearm, and the individual was wiggling their thumb through the mud.<ref name=Zhang2021/> | |||
| == Interbreeding == | |||
| {{see also|Interbreeding between archaic and modern humans}} | |||
| Analyses of the modern human genomes indicate past interbreeding with at least two groups of archaic humans, Neanderthals<ref name=Pennisi>{{cite journal |doi=10.1126/science.340.6134.799 |pmid=23687020 |title=More Genomes from Denisova Cave Show Mixing of Early Human Groups |journal=] |volume=340 |issue=6134 |pages=799 |year=2013 |last1=Pennisi |first1=E. |author-link=Elizabeth Pennisi |bibcode=2013Sci...340..799P}}</ref> and Denisovans,<ref name=pmid21179161/><ref>{{cite journal |vauthors=Green RE, Krause J, Briggs AW |title=A draft sequence of the Neandertal genome |journal=] |volume=328 |issue=5979 |pages=710–22 |year=2010 |pmid=20448178 |pmc=5100745 |doi=10.1126/science.1188021 |url=http://www.eva.mpg.de/neandertal/press/presskit-neandertal/pdf/Science_Green.pdf |display-authors=etal |bibcode=2010Sci...328..710G |access-date=3 May 2013 |archive-date=13 August 2012 |archive-url=https://web.archive.org/web/20120813021203/http://www.eva.mpg.de/neandertal/press/presskit-neandertal/pdf/Science_Green.pdf |url-status=live}}</ref> and that such interbreeding events occurred on multiple occasions. Comparisons of the Denisovan, Neanderthal, and modern human genomes have revealed evidence of a complex web of interbreeding among these lineages.<ref name=Pennisi /> | |||
| === Archaic humans === | |||
| As much as 17% of the Denisovan genome from Denisova Cave represents DNA from the local Neanderthal population.<ref name=Pennisi /> Denisova 11 was an ] (first generation) Denisovan/Neanderthal hybrid; the fact that such an individual was found may indicate interbreeding was a common occurrence here.<ref name="NAT-20180822">{{cite journal |last=Warren |first=M. |title=Mum's a Neanderthal, Dad's a Denisovan: First discovery of an ancient-human hybrid – Genetic analysis uncovers a direct descendant of two different groups of early humans. |year= 2018 |journal=] |volume=560 |issue=7719 |pages=417–418 |doi=10.1038/d41586-018-06004-0 |pmid=30135540 |bibcode=2018Natur.560..417W |doi-access=free}}</ref> The Denisovan genome shares more derived alleles with the Altai Neanderthal genome from Siberia than with the ] Neanderthal genome from Croatia or the ] Neanderthal genome from the Caucasus, suggesting that the gene flow came from a population that was more closely related to the local Altai Neanderthals.<ref name=prualtai/> However, Denny's Denisovan father had the typical Altai Neanderthal ], while her Neanderthal mother represented a population more closely related to Vindija Neanderthals.<ref>{{cite journal|last=Warren |first=Matthew |title=Mum's a Neanderthal, Dad's a Denisovan: First discovery of an ancient-human hybrid |journal=] |volume=560 |issue=7719 |pages=417–18 |year=2018 |doi=10.1038/d41586-018-06004-0 |pmid=30135540 |bibcode=2018Natur.560..417W |doi-access=free}}</ref> Denisova 25, dated to 200ka, is estimated to have inherited 5% of his genome from a previously unknown population of Neanderthals, and came from a different population of Denisovans than the younger samples.<ref name="h523"/> | |||
| About 4% of the Denisovan genome derives from an unidentified archaic hominin,<ref name=Pennisi /> perhaps the source of the anomalous ancient mtDNA, indicating this species diverged from Neanderthals and humans over a million years ago. The only identified ''Homo'' species of ] Asia are ''H. erectus'' and ''H. heidelbergensis'',<ref name=prualtai>{{cite journal|last1=Prüfer |first1=K. |last2=Racimo |first2=Fernando |last3=Patterson |first3=N. |last4=Jay |first4=F. |last5=Sankararaman |first5=S. |last6=Sawyer |first6=S. |title=The complete genome sequence of a Neanderthal from the Altai Mountains |journal=] |year=2013 |volume=505 |issue=7481 |pages=43–49 |doi=10.1038/nature12886 |display-authors=et al. |bibcode=2014Natur.505...43P |pmid=24352235 |pmc=4031459}}</ref><ref>{{cite journal|first1=A. B. |last1=Wolf |first2=J. M. |last2=Akey |year=2018 |title=Outstanding questions in the study of archaic hominin admixture |journal=] |volume=14 |issue=5 |page=e1007349 |doi=10.1371/journal.pgen.1007349 |pmc=5978786 |pmid=29852022 |doi-access=free }}</ref> though in 2021, specimens allocated to the latter species were reclassified as '']'' and '']''.<ref name=Ni2021>{{cite journal |first1=X. |last1=Ni |first2=Q. |last2=Ji |first3=W. |last3=Wu |display-authors=et al. |year=2021 |title=Massive cranium from Harbin in northeastern China establishes a new Middle Pleistocene human lineage |journal=Innovation |volume=2 |issue=3 |page=100130 |doi=10.1016/j.xinn.2021.100130 |issn=2666-6758 |pmid=34557770 |pmc=8454562 |bibcode=2021Innov...200130N |s2cid=236784246}}</ref> | |||
| Before splitting from Neanderthals, their ancestors ("Neandersovans") migrating into Europe apparently interbred with an unidentified "superarchaic" human species who were already present there; these superarchaics were the descendants of a very early migration out of Africa around 1.9 mya.<ref>{{cite journal |first1=A. R. |last1=Rogers |first2=N. S. |last2=Harris |first3=A. A. |last3=Achenbach |year=2020 |title=Neanderthal-Denisovan ancestors interbred with a distantly related hominin |journal=] |volume=6 |issue=8 |page=eaay5483 |doi=10.1126/sciadv.aay5483 |pmid=32128408 |pmc=7032934 |bibcode=2020SciA....6.5483R}}</ref> | |||
| === Modern humans === | |||
| {{Human timeline}} | |||
| A 2011 study found that Denisovan DNA is prevalent in ], ], ]ns, ], ], Eastern ], and ] (from the Philippines); but not in ], western Indonesians, ] (from Malaysia), or ] (from the ]). This may suggest that Denisovan introgression occurred within the Pacific region rather than on the Asian mainland, and that ancestors of the latter groups were not present in Southeast Asia at the time.<ref name="Reich">{{cite journal |doi=10.1016/j.ajhg.2011.09.005 |pmid=21944045 |pmc=3188841 |title=Denisova Admixture and the First Modern Human Dispersals into Southeast Asia and Oceania |journal=The American Journal of Human Genetics |volume=89 |issue=4 |pages=516–28 |year=2011 |last1=Reich |first1=David |last2=Patterson |first2=Nick |last3=Kircher |first3=Martin |last4=Delfin |first4=Frederick |last5=Nandineni |first5=Madhusudan R |last6=Pugach |first6=Irina |last7=Ko |first7=Albert Min-Shan |last8=Ko |first8=Ying-Chin |last9=Jinam |first9=Timothy A. |last10=Phipps |first10=Maude E |last11=Saitou |first11=Naruya |last12=Wollstein |first12=Andreas |last13=Kayser |first13=Manfred |last14=Pääbo |first14=Svante |author14-link=Svante Pääbo |last15=Stoneking |first15=Mark |display-authors=4}}</ref><ref>{{Cite journal |last=Yang |first=Melinda A. |date=2022-01-06 |title=A genetic history of migration, diversification, and admixture in Asia |url=https://www.pivotscipub.com/hpgg/2/1/0001 |journal=Human Population Genetics and Genomics |language=en |volume=2 |issue=1 |pages=1–32 |doi=10.47248/hpgg2202010001 |issn=2770-5005 |doi-access=free |access-date=5 December 2023 |archive-date=16 April 2023 |archive-url=https://web.archive.org/web/20230416014152/https://www.pivotscipub.com/hpgg/2/1/0001 |url-status=live }}</ref> In the ] genome, about 4–6%<ref name=pmid21179161/> or 1.9–3.4% derives from Denisovan introgression.<ref>{{cite journal|first=B. |last=Vernot |display-authors=et al. |year=2016 |title=Excavating Neandertal and Denisovan DNA from the genomes of Melanesian individuals |journal=] |volume=352 |issue=6282 |pages=235–239 |doi=10.1126/science.aad9416 |pmid=26989198 |pmc=6743480 |bibcode=2016Sci...352..235V}}</ref> Prior to 2021, New Guineans and Australian Aborigines were reported to have the most introgressed DNA,<ref name="Cooper"/> but Australians have less than New Guineans.<ref name=Rasmussen2011>{{cite journal |last1=Rasmussen |first1=M. |first2=Xiaosen |last2=Guo |first3=Yong |last3=Wang |first4=Kirk E. |last4=Lohmueller |first5=Simon |last5=Rasmussen |display-authors=et al. |year=2011 |title=An Aboriginal Australian genome reveals separate human dispersals into Asia |journal=] |volume=334 |issue=6052 |pages=94–98 |doi=10.1126/science.1211177 |pmid=21940856 |pmc=3991479 |bibcode=2011Sci...334...94R}}</ref> A 2021 study discovered 30 to 40% more Denisovan ancestry in ] in the Philippines than in ], estimated as about 5% of the genome. The Aeta Magbukon in Luzon have the highest known proportion of Denisovan ancestry of any population in the world.<ref name=Larena2021>{{cite journal |last1=Larena |first1=Maximilian |last2=McKenna |first2=James |last3=Sanchez-Quinto |first3=Federico |last4=Bernhardsson |first4=Carolina |last5=Ebeo |first5=Carlo |last6=Reyes |first6=Rebecca |last7=Casel |first7=Ophelia |last8=Huang |first8=Jin-Yuan |last9=Hagada |first9=Kim Pullupul |last10=Guilay |first10=Dennis |last11=Reyes |first11=Jennelyn |last12=Allian |first12=Fatima Pir |last13=Mori |first13=Virgilio |last14=Azarcon |first14=Lahaina Sue |last15=Manera |first15=Alma |last16=Terando |first16=Celito |last17=Jamero |first17=Lucio |last18=Sireg |first18=Gauden |last19=Manginsay-Tremedal |first19=Renefe |last20=Labos |first20=Maria Shiela |last21=Vilar |first21=Richard Dian |last22=Latiph |first22=Acram |last23=Saway |first23=Rodelio Linsahay |last24=Marte |first24=Erwin |last25=Magbanua |first25=Pablito |last26=Morales |first26=Amor |last27=Java |first27=Ismael |last28=Reveche |first28=Rudy |last29=Barrios |first29=Becky |last30=Burton |first30=Erlinda |last31=Salon |first31=Jesus Christopher |last32=Kels |first32=Ma. Junaliah Tuazon |last33=Albano |first33=Adrian |last34=Cruz-Angeles |first34=Rose Beatrix |last35=Molanida |first35=Edison |last36=Granehäll |first36=Lena |last37=Vicente |first37=Mário |last38=Edlund |first38=Hanna |last39=Loo |first39=Jun-Hun |last40=Trejaut |first40=Jean |last41=Ho |first41=Simon Y.W. |last42=Reid |first42=Lawrence |last43=Lambeck |first43=Kurt |last44=Malmström |first44=Helena |last45=Schlebusch |first45=Carina |last46=Endicott |first46=Phillip |last47=Jakobsson |first47=Mattias |display-authors=4 |title=Philippine Ayta possess the highest level of Denisovan ancestry in the world |journal=] |date=August 2021 |volume=31 |issue=19 |pages=4219–4230.e10 |doi=10.1016/j.cub.2021.07.022 |pmid=34388371 |pmc=8596304 |s2cid=236994320 |doi-access=free|bibcode=2021CBio...31E4219L }}</ref> In Papuans, less Denisovan ancestry is seen in the ] than ], and some autosomes (such as ]) also have less Denisovan ancestry, which could indicate ]. The former observation could also be explained by less female Denisovan introgression into modern humans, or more female modern human immigrants who diluted Denisovan X chromosome ancestry.<ref name=Meyer2012/> | |||
| In contrast, 0.2% derives from Denisovan ancestry in mainland Asians and ]s.<ref name="NAT-2013">{{cite journal |doi=10.1038/nature12886 |pmid=24352235 |pmc=4031459 |title=The complete genome sequence of a Neanderthal from the Altai Mountains |journal=] |volume=505 |issue=7481 |pages=43–49 |year=2013 |last1=Prüfer |first1=K. |last2=Racimo |first2=F. |last3=Patterson |first3=N.|last4=Jay |first4=F.|last5=Sankararaman |first5=S. |last6=Sawyer |first6=S. |last7=Heinze |first7=A.|last8=Renaud |first8=G. |last9=Sudmant |first9=P. H. |last10=De Filippo |first10=C. |display-authors=4 |bibcode=2014Natur.505...43P}}</ref> ] were found to have levels of Denisovan admixture similar to that seen in East Asians.<ref name="Browning Browning Zhou Tucci 2018"/> The discovery of the 40,000-year-old Chinese modern human ] lacking Denisovan DNA significantly different from the levels in modern-day East Asians discounts the hypothesis that immigrating modern humans simply diluted Denisovan ancestry whereas Melanesians lived in reproductive isolation.<ref>{{cite journal|first1=Q. |last1=Fu |first2=M. |last2=Meyer |first3=X. |last3=Gao |display-authors=et al. |year=2013 |title=DNA analysis of an early modern human from Tianyuan Cave, China |journal=] |volume=110 |issue=6 |pages=2223–2227 |doi=10.1073/pnas.1221359110 |pmid=23341637 |pmc=3568306 |bibcode=2013PNAS..110.2223F |doi-access=free}}</ref><ref name="Cooper"/> A 2018 study of Han Chinese, ], and ] genomes showed that modern East Asians have DNA from two different Denisovan populations: one similar to the Denisovan DNA found in Papuan genomes, and a second that is closer to the Denisovan genome from Denisova Cave. This could indicate two separate introgression events involving two different Denisovan populations. In South Asian genomes, DNA only came from the same single Denisovan introgression seen in Papuans.<ref name="Browning Browning Zhou Tucci 2018">{{cite journal |last1=Browning |first1=S. R. |author1-link=Sharon R. Browning |last2=Browning |first2=B. L. |last3=Zhou |first3=Yi. |last4=Tucci |first4=S. |last5=Akey |first5=J. M. |display-authors=4 |title=Analysis of Human Sequence Data Reveals Two Pulses of Archaic Denisovan Admixture |journal=] |volume=173 |issue=1 |pages=53–61.e9 |year=2018 |issn=0092-8674 |url= |doi=10.1016/j.cell.2018.02.031 |pmid=29551270 |pmc=5866234}}</ref> A 2019 study found a third wave of Denisovans which introgressed into East Asians. Introgression, also, may not have immediately occurred when modern humans immigrated into the region.<ref name=JacobsG2019/> | |||
| The timing of introgression into Oceanian populations likely occurred after Eurasians and Oceanians split roughly 58,000 years ago, and before Papuan and Aboriginal Australians split from each other roughly 37,000 years ago. Given the present day distribution of Denisovan DNA, this may have taken place in Wallacea, though the discovery of a 7,200 year old ] girl (closely related to Papuans and Aboriginal Australians) from Sulawesi carrying Denisovan DNA makes Sundaland another potential candidate. Other early Sunda hunter gatherers so far sequenced carry very little Denisovan DNA, which either means the introgression event did not take place in Sundaland, or Denisovan ancestry was diluted with gene flow from the mainland Asian ] culture and subsequent ] cultures.<ref>{{Cite journal|first=Selina |last=Carlhoff |title=Genome of a middle Holocene hunter-gatherer from Wallace |year=2021 |journal=] |volume=596 |issue=7873 |pages=543–547 |doi=10.1038/s41586-021-03823-6 |pmid=34433944 |pmc=8387238 |bibcode=2021Natur.596..543C}}</ref> | |||
| In other regions of the world, archaic introgression into humans stems from a group of Neanderthals related to those which inhabited ], Croatia, as opposed to archaics related to Siberian Neanderthals and Denisovans. However, about 3.3% of the archaic DNA in the modern Icelandic genome descends from the Denisovans, and such a high percentage could indicate a western Eurasian population of Denisovans which introgressed into either Vindija-related Neanderthals or immigrating modern humans.<ref name=Skov2020>{{cite journal|first1=L. |last1=Skov |first2=M. C. |last2=Macià |first3=G. |last3=Sveinbjörnsson |display-authors=et al. |year=2020 |title=The nature of Neanderthal introgression revealed by 27,566 Icelandic genomes |journal=] |volume=582 |issue=7810 |pages=78–83 |doi=10.1038/s41586-020-2225-9 |pmid=32494067 |bibcode=2020Natur.582...78S |s2cid=216076889}}</ref> | |||
| Denisovan genes may have helped early modern humans migrating out of Africa to acclimatize{{Citation needed|date=June 2023}}. Although not present in the sequenced Denisovan genome, the distribution pattern and divergence of HLA-B*73 from other ] alleles (involved in the ]'s ]) has led to the suggestion that it introgressed from Denisovans into modern humans in ]. In a 2011 study, half of the HLA alleles of modern Eurasians were shown to represent archaic HLA haplotypes, and were inferred to be of Denisovan or Neanderthal origin.<ref name="10.1126/science.1209202">{{cite journal |doi=10.1126/science.1209202 |pmid=21868630 |pmc=3677943 |title=The Shaping of Modern Human Immune Systems by Multiregional Admixture with Archaic Humans |journal=] |volume=334 |issue=6052 |pages=89–94 |year=2011 |last1=Abi-Rached |first1=L. |last2=Jobin |first2=M. J. |last3=Kulkarni |first3=S. |last4=McWhinnie |first4=A. |last5=Dalva |first5=K. |last6=Gragert |first6=L. |last7=Babrzadeh |first7=F. |last8=Gharizadeh |first8=B. |last9=Luo |first9=M. |last10=Plummer |first10=F. A. |last11=Kimani |first11=J. |last12=Carrington |first12=M. |last13=Middleton |first13=D. |last14=Rajalingam |first14=R. |last15=Beksac |first15=M. |last16=Marsh |first16=S. G. E. |last17=Maiers |first17=M. |last18=Guethlein |first18=L. A. |last19=Tavoularis |first19=S. |last20=Little |first20=A.-M. |last21=Green |first21=R. E. |last22=Norman |first22=P. J. |last23=Parham |first23=P. |display-authors=4 |bibcode=2011Sci...334...89A}}</ref> A haplotype of EPAS1 in modern Tibetans, which allows them to live at high elevations in a low-oxygen environment, likely came from Denisovans.<ref name=Huerta2014/><ref name=ChenBaishiya /> Genes related to ] transporters (which are involved in ]) and to ]s (involved in smelling) are more active in people with more Denisovan ancestry.<ref name=Sankararaman2016>{{cite journal|first1=S. |last1=Sankararaman |first2=S. |last2=Mallick |first3=N. |last3=Patterson |first4=D. |last4=Reich |year=2016 |title=The combined landscape of Denisovan and Neanderthal ancestry in present-day humans |journal=] |volume=26 |issue=9 |pages=1241–1247 |doi=10.1016/j.cub.2016.03.037 |pmc=4864120 |pmid=27032491|bibcode=2016CBio...26.1241S }}</ref> Denisovan genes may have conferred a degree of immunity against the G614 mutation of ].<ref>{{Cite journal|last=Muscat |first=Baron Y. |date=2021 |title=Could the Denisovan Genes have conferred enhanced Immunity Against the G614 Mutation of SARS-CoV-2? |url=https://www.researchgate.net/publication/352888303 |journal=] |volume=36 |access-date=19 July 2021 |archive-date=19 July 2021 |archive-url=https://web.archive.org/web/20210719232527/https://www.researchgate.net/profile/Yves-Muscat-Baron/publication/352888303_Muscat_Baron_Y_Could_the_Denisovan_Genes_have_conferred_enhanced_Immunity_against_the_G614_Mutation_of_SARS-CoV-2/links/60de0c24a6fdccb745fbaa51/Muscat-Baron-Y-Could-the-Denisovan-Genes-have-conferred-enhanced-Immunity-against-the-G614-Mutation-of-SARS-CoV-2.pdf |url-status=live}}</ref> Denisovan introgressions may have influenced the immune system of present-day Papuans and potentially favoured "variants to immune-related phenotypes" and "adaptation to the local environment".<ref>{{Cite journal |last1=Vespasiani |first1=Davide M. |last2=Jacobs |first2=Guy S. |last3=Cook |first3=Laura E. |last4=Brucato |first4=Nicolas |last5=Leavesley |first5=Matthew |last6=Kinipi |first6=Christopher |last7=Ricaut |first7=François-Xavier |last8=Cox |first8=Murray P. |last9=Gallego Romero |first9=Irene |date=2022-12-08 |title=Denisovan introgression has shaped the immune system of present-day Papuans |journal=PLOS Genetics |volume=18 |issue=12 |pages=e1010470 |doi=10.1371/journal.pgen.1010470 |issn=1553-7390 |pmc=9731433 |pmid=36480515 |doi-access=free }}</ref> | |||
| In December 2023, scientists reported that ]s inherited by ]s from ]s and Denisovans may biologically influence the daily routine of modern humans.<ref name="NYT-20231214">{{cite news |last=Zimmer |first=Carl |authorlink=Carl Zimmer |title=Morning Person? You Might Have Neanderthal Genes to Thank. - Hundreds of genetic variants carried by Neanderthals and Denisovans are shared by people who like to get up early. |url=https://www.nytimes.com/2023/12/14/science/neanderthal-sleep-morning-people.html |date=14 December 2023 |work=] |url-status=live |archiveurl=https://archive.today/20231214070730/https://www.nytimes.com/2023/12/14/science/neanderthal-sleep-morning-people.html |archivedate=14 December 2023 |accessdate=14 December 2023 }}</ref> | |||
| == See also == | |||
| {{Portal|Biology}} | |||
| * {{annotated link|Red Deer Cave people}} | |||
| * {{annotated link|Homo longi|''Homo longi''}} | |||
| * {{annotated link|Homo naledi|''Homo naledi''}} | |||
| * {{annotated link|Nesher Ramla Homo|Nesher Ramla ''Homo''}} | |||
| * {{annotated link|Timeline of human evolution}} | |||
| == References == | |||
| {{reflist}} | {{reflist}} | ||
| ==Further reading== | == Further reading == | ||
| * {{cite thesis |last1=Karlsson |first1=Mattis |title=From Fossil To Fact: The Denisova Discovery as Science in Action |publisher=LiU E-press |isbn=9789179291716 |url=http://www.diva-portal.org/smash/record.jsf?pid=diva2%3A1632719&dswid=-2272 |access-date=18 March 2022}} | |||
| * {{Citation | |||
| | url = http://www.newscientist.com/article/mg21128254.000-stone-age-toe-could-redraw-human-family-tree.html | |||
| | title = Stone Age toe could redraw human family tree | |||
| | work = ] | |||
| | date = 13 August 2011 | |||
| | first = Colin | |||
| | last = Barras}}. | |||
| * {{Citation | |||
| | url = http://www.reuters.com/article/idUSTRE62N4VS20100324 | |||
| | title = Possible new human ancestor found in Siberia | |||
| | work = ] | |||
| | date = 24 March 2010 | |||
| | first = Maggie | |||
| | last = Fox}}. | |||
| * {{Citation | |||
| | url = http://www.bbc.co.uk/news/science-environment-12059564 | |||
| | title = Ancient humans, dubbed 'Denisovans', interbred with us | |||
| | publisher = ] | |||
| | date = 22 December 2010 | |||
| | quote = The study shows that Denisovans interbred with the ancestors of the present day people of the Melanesian region north and north-east of Australia. Melanesian DNA comprises between 4% and 6% Denisovan DNA. | |||
| | first = Pallab | |||
| | last = Ghosh }} | |||
| * {{Citation | |||
| | url = http://johnhawks.net/weblog/reviews/neandertals/neandertal_dna/denisova-nuclear-genome-reich-2010.html | |||
| | title = The Denisova genome FAQ | |||
| | work = ] | |||
| | date = 22 December 2010 | |||
| | first = John | |||
| | last = Hawks}}. | |||
| * {{Citation | |||
| | url = http://news.bbc.co.uk/1/hi/sci/tech/8583254.stm | |||
| | title = DNA identifies new ancient human dubbed 'X-woman' | |||
| | publisher = ] | |||
| | date = 25 March 2010 | |||
| | first = Paul | |||
| | last = Rincon}}. | |||
| * {{Citation | |||
| | url = http://ngm.nationalgeographic.com/2013/07/125-missing-human-ancestor/shreeve-text | |||
| | title = The Case of the Missing Ancestor | |||
| | newspaper = ] | |||
| | date = July 2013 | |||
| | first = Jamie | |||
| | last = Shreeve }}. | |||
| * {{Citation | |||
| | url = http://www.nytimes.com/2010/03/25/science/25human.html | |||
| | title = Bone May Reveal a New Human Group | |||
| | newspaper = ] | |||
| | date = 24 March 2010 | |||
| | first = Nicholas | |||
| | last = Wade }}. | |||
| * {{Citation | |||
| | url = http://blogs.discovermagazine.com/loom/2010/03/24/the-x-womans-fingerbone/ | |||
| | title = Hybrid speculation (blog) | |||
| | publisher = ] | |||
| | date = 24 March 2010 | |||
| | first = Carl | |||
| | last = Zimmer }}. | |||
| * {{Citation | |||
| | url = http://blogs.discovermagazine.com/loom/2010/12/22/meet-the-denisovans-the-newest-members-of-the-human-tree-of-life/ | |||
| | title = Meet the Denisovans, the newest members of the human tree of life (blog) | |||
| | publisher = ] | |||
| | date = 22 December 2010 | |||
| | first = Carl | |||
| | last = Zimmer }}. | |||
| ==External links== | == External links == | ||
| * {{Commons category-inline}} | |||
| * | * | ||
| * – ], ] (August 2016). | |||
| * | |||
| {{Human Evolution}} | {{Human Evolution}} | ||
| {{Prehistoric technology}} | {{Prehistoric technology}} | ||
| ] | ] | ||
| ] | ] | ||
| ] | |||
| ] | |||
| ] | ] | ||
| ] | ] | ||
| ] | |||
| ] | ] | ||
| ] | |||
| {{Link FA|de}} | |||
Latest revision as of 20:24, 10 January 2025
Asian archaic human

The Denisovans or Denisova hominins (/dəˈniːsəvə/ də-NEE-sə-və) are an extinct species or subspecies of archaic human that ranged across Asia during the Lower and Middle Paleolithic, and lived, based on current evidence, from 285 to 25 thousand years ago. Denisovans are known from few physical remains; consequently, most of what is known about them comes from DNA evidence. No formal species name has been established pending more complete fossil material.
The first identification of a Denisovan individual occurred in 2010, based on mitochondrial DNA (mtDNA) extracted from a juvenile female finger bone excavated from the Siberian Denisova Cave in the Altai Mountains in 2008. Nuclear DNA indicates close affinities with Neanderthals. The cave was also periodically inhabited by Neanderthals, but it is unclear whether Neanderthals and Denisovans ever cohabited in the cave. Additional specimens from Denisova Cave were subsequently identified, as was a single specimen from the Baishiya Karst Cave on the Tibetan Plateau, and Cobra Cave in the Annamite Mountains of Laos. DNA evidence suggests they had dark skin, eyes, and hair, and had a Neanderthal-like build and facial features. However, they had larger molars which are reminiscent of Middle to Late Pleistocene archaic humans and australopithecines.
Denisovans apparently interbred with modern humans, with a high percentage (roughly 5%) occurring in Melanesians, Aboriginal Australians, and Filipino Negritos. This distribution suggests that there were Denisovan populations across Asia. There is also evidence of interbreeding with the Altai Neanderthal population, with about 17% of the Denisovan genome from Denisova Cave deriving from them. A first-generation hybrid nicknamed "Denny" was discovered with a Denisovan father and a Neanderthal mother. Additionally, 4% of the Denisovan genome comes from an unknown archaic human species, which diverged from modern humans over one million years ago.
Taxonomy
Denisovans may represent a new species of Homo or an archaic subspecies of Homo sapiens (modern humans), but there are too few fossils to erect a proper taxon. Proactively proposed species names for Denisovans are H. denisova or H. altaiensis. Chinese researchers suggest the Denisovans were members of Homo longi, and the idea has been supported by the palaeontologist Chris Stringer. In 2024, paleoanthropologists Christopher Bae and Xiujie Wu designated the Xujiayao hominin fossils as the holotype of the species Homo juluensis, and suggested sinking Denisovans into this species.
Discovery

Denisova Cave is located in Altai Krai, Russia, in south-central Siberia, on the western edges of the Altai Mountains. It is named after Denis (Dyonisiy), a Russian Old Believer hermit who lived there in the 18th century. The cave was first inspected for fossils in the 1970s by Soviet paleontologist Nikolai Ovodov, who was looking for remains of canids.
In 2008, Michael Shunkov from the Russian Academy of Sciences and other Russian archaeologists from the Institute of Archaeology and Ethnography of the Siberian Branch of the Russian Academy of Sciences in Novosibirsk Akademgorodok investigated the cave and found the finger bone of a juvenile female hominin originally dated to 50–30,000 years ago. The estimate has changed to 76,200–51,600 years ago. The specimen was originally named X-woman because matrilineal mitochondrial DNA (mtDNA) extracted from the bone demonstrated it to belong to a novel ancient hominin, genetically distinct both from contemporary modern humans and from Neanderthals.
In 2019, Greek archaeologist Katerina Douka and colleagues radiocarbon dated specimens from Denisova Cave, and estimated that Denisova 2 (the oldest specimen) lived 195,000–122,700 years ago. Older Denisovan DNA collected from sediments in the East Chamber dates to 217,000 years ago. Based on artifacts also discovered in the cave, hominin occupation (most likely by Denisovans) began 287±41 or 203±14 ka. Neanderthals were also present 193±12 ka and 97±11 ka, possibly concurrently with Denisovans.
Specimens
"Woman X" redirects here. For other uses, see X woman (disambiguation).The fossils of multiple distinct Denisovan individuals from Denisova Cave have been identified through their ancient DNA (aDNA): Denisova 2, 3, 4, 8, 11, and 25. An mtDNA-based phylogenetic analysis of these individuals suggested that Denisova 2 is the oldest, followed by Denisova 8, while Denisova 3 and Denisova 4 were roughly contemporaneous. In 2024, scientists announced the sequence of Denisova 25, which was in a layer dated to 200ka. During DNA sequencing, a low proportion of the Denisova 2, Denisova 4 and Denisova 8 genomes were found to have survived, but a high proportion of the Denisova 3 and Denisova 25 genomes were intact. The Denisova 3 sample was cut into two, and the initial DNA sequencing of one fragment was later independently confirmed by sequencing the mtDNA from the second.
Denisova Cave contained the only known examples of Denisovans until 2019, when a research group led by Fahu Chen, Dongju Zhang, and Jean-Jacques Hublin described a partial mandible discovered in 1980 by a Buddhist monk in the Baishiya Karst Cave on the Tibetan Plateau in China. Known as the Xiahe mandible, the fossil became part of the collection of Lanzhou University, where it remained unstudied until 2010. It was determined by ancient protein analysis to contain collagen that by sequence was found to have close affiliation to that of the Denisovans from Denisova Cave, while uranium decay dating of the carbonate crust enshrouding the specimen indicated it was more than 160,000 years old. The identity of this population was later confirmed through study of environmental DNA, which found Denisovan mtDNA in sediment layers ranging in date from 100,000 to 60,000 years before present, and perhaps more recent. A 2024 reanalysis identified a partial Denisovan rib fragment dating to between 48,000 BP and 32,000 BP.
In 2018, a team of Laotian, French, and American anthropologists, who had been excavating caves in the Laotian jungle of the Annamite Mountains since 2008, was directed by local children to the site Tam Ngu Hao 2 ("Cobra Cave") where they recovered a human tooth. The tooth (catalogue number TNH2-1) developmentally matches a 3.5 to 8.5 year old, and a lack of amelogenin (a protein on the Y chromosome) suggests it belonged to a girl barring extreme degradation of the protein over a long period of time. Dental proteome analysis was inconclusive for this specimen, but the team found it anatomically comparable with the Xiahe mandible, and so tentatively categorized it as a Denisovan, although they could not rule out it being Neanderthal. The tooth probably dates to 164,000 to 131,000 years ago.
Some older findings may or may not belong to the Denisovan line, but Asia is not well mapped in regards to human evolution. Such findings include the Dali skull, the Xujiayao hominin, Maba Man, the Jinniushan hominin, and the Narmada Human. The Xiahe mandible shows morphological similarities to some later East Asian fossils such as Penghu 1, but also to Chinese H. erectus. In 2021, Chinese palaeoanthropologist Qiang Ji suggested his newly erected species, H. longi, may represent the Denisovans based on the similarity between the type specimen's molar and that of the Xiahe mandible. In 2024, Bae and Wu suggested classifying the Xujiayao and Denisovan material as H. juluensis, and the Dali Man and similar specimens as H. longi.
| Name | Fossil elements | Age | Discovery | Place | Sex and age | Publication | Image | GenBank accession |
|---|---|---|---|---|---|---|---|---|
| Denisova 3 (also known as X Woman) |
Distal phalanx of the fifth finger | 76.2–51.6 ka | 2008 | Denisova cave (Russia) | 13.5-year-old adolescent female | 2010 |  |
NC013993 |
| Denisova 4 | Permanent upper 2nd or 3rd molar | 84.1–55.2 ka | 2000 | Denisova cave (Russia) | Adult male | 2010 |  |
FR695060 |
| Denisova 8 | Permanent upper 3rd molar | 136.4–105.6 ka | 2010 | Denisova cave (Russia) | Adult male | 2015 | KT780370 | |
| Denisova 2 | Deciduous 2nd lower molar | 194.4–122.7 ka | 1984 | Denisova cave (Russia) | Adolescent female | 2017 | KX663333 | |
| Xiahe mandible | Partial mandible | > 160 ka | 1980 | Baishiya Cave (China) | 2019 | 
|
||
| Denisova 11 (also known as Denny, Denisovan x Neanderthal hybrid) |
Arm or leg bone fragment | 118.1–79.3 ka | 2012 | Denisova cave (Russia) | 13 year old adolescent female | 2016 | 
|
|
| Denisova 13 | Parietal bone fragment | Found in layer 22 which dates to ~285±39 ka | 2019 | Denisova cave (Russia) | pending | |||
| TNH2-1 | Permanent lower left 1st or 2nd molar | 164–131 ka | 2018 | Tam Ngu Hao 2 cave (Laos) | 3.5 to 8.5 year old female | 2022 | 
|
|
| BSY-19-B896-1 (Xiahe 2) | Distal rib fragment | 48–32 ka | 1980 | Baishiya Cave (China) | Unknown | 2024 | 
| |
| Denisova 25 | Molar | 200 ka | 2024 | Denisova cave (Russia) | Male | pending |
Evolution
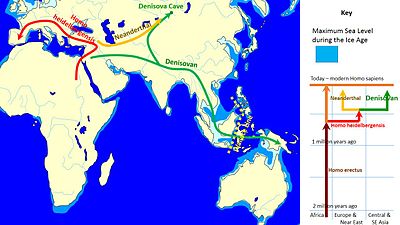
Sequenced mitochondrial DNA (mtDNA), preserved by the cool climate of the cave (average temperature is at freezing point), was extracted from Denisova 3 by a team of scientists led by Johannes Krause and Svante Pääbo from the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany. Denisova 3's mtDNA differs from that of modern humans by 385 bases (nucleotides) out of approximately 16,500, whereas the difference between modern humans and Neanderthals is around 202 bases. In comparison, the difference between chimpanzees and modern humans is approximately 1,462 mtDNA base pairs. This suggested that Denisovan mtDNA diverged from that of modern humans and Neanderthals about 1,313,500–779,300 years ago; whereas modern human and Neanderthal mtDNA diverged 618,000–321,200 years ago. Krause and colleagues then concluded that Denisovans were the descendants of an earlier migration of H. erectus out of Africa, completely distinct from modern humans and Neanderthals.
However, according to the nuclear DNA (nDNA) of Denisova 3—which had an unusual degree of DNA preservation with only low-level contamination—Denisovans and Neanderthals were more closely related to each other than they were to modern humans. Using the percent distance from human–chimpanzee last common ancestor, Denisovans/Neanderthals split from modern humans about 804,000 years ago, and from each other 640,000 years ago. Using a mutation rate of 1×10 or 0.5×10 per base pair (bp) per year, the Neanderthal/Denisovan split occurred around either 236–190,000 or 473–381,000 years ago respectively. Using 1.1×10 per generation with a new generation every 29 years, the time is 744,000 years ago. Using 5×10 nucleotide site per year, it is 616,000 years ago. Using the latter dates, the split had likely already occurred by the time hominins spread out across Europe. H. heidelbergensis is typically considered to have been the direct ancestor of Denisovans and Neanderthals, and sometimes also modern humans. Due to the strong divergence in dental anatomy, they may have split before characteristic Neanderthal dentition evolved about 300,000 years ago.
The more divergent Denisovan mtDNA has been interpreted as evidence of admixture between Denisovans and an unknown archaic human population, possibly a relict H. erectus or H. erectus-like population about 53,000 years ago. Alternatively, divergent mtDNA could have also resulted from the persistence of an ancient mtDNA lineage which only went extinct in modern humans and Neanderthals through genetic drift. Modern humans contributed mtDNA to the Neanderthal lineage, but not to the Denisovan mitochondrial genomes yet sequenced. The mtDNA sequence from the femur of a 400,000-year-old H. heidelbergensis from the Sima de los Huesos Cave in Spain was found to be related to those of Neanderthals and Denisovans, but closer to Denisovans, and the authors posited that this mtDNA represents an archaic sequence which was subsequently lost in Neanderthals due to replacement by a modern-human-related sequence.
Demographics
See also: Archaic humans in Southeast Asia
Denisovans are known to have lived in Siberia, Tibet, and Laos. The Xiahe mandible is the earliest recorded human presence on the Tibetan Plateau. Though their remains have been identified in only these three locations, traces of Denisovan DNA in modern humans suggest they ranged across East Asia, and potentially western Eurasia. In 2019, geneticist Guy Jacobs and colleagues identified three distinct populations of Denisovans responsible for the introgression into modern populations now native to, respectively: Siberia and East Asia; New Guinea and nearby islands; and Oceania and, to a lesser extent, across Asia. Using coalescent modeling, the Denisova Cave Denisovans split from the second population about 283,000 years ago; and from the third population about 363,000 years ago. This indicates that there was considerable reproductive isolation between Denisovan populations. In a 2024 study, scientist Danat Yermakovich, of the University of Tartu, discovered that people living at different elevations in Papua New Guinea have differences in Denisovan DNA; with people living in the highlands having variants for early brain development and those living in the lowlands having variants for the immune system.
Based on the high percentages of Denisovan DNA in modern Papuans and Australians, Denisovans may have crossed the Wallace Line into these regions (with little back-migration west), the second known human species to do so, along with earlier Homo floresiensis. By this logic, they may have also entered the Philippines, living alongside H. luzonensis which, if this is the case, may represent the same or a closely related species. These Denisovans may have needed to cross large bodies of water. Alternately, high Denisovan DNA admixture in modern Papuan populations may simply represent higher mixing among the original ancestors of Papuans prior to crossing the Wallace line. Icelanders also have an anomalously high Denisovan heritage, which could have stemmed from a Denisovan population far west of the Altai Mountains. Genetic data suggests Neanderthals were frequently making long crossings between Europe and the Altai Mountains especially towards the date of their extinction.
Using exponential distribution analysis on haplotype lengths, Jacobs calculated introgression into modern humans occurred about 29,900 years ago with the Denisovan population ancestral to New Guineans; and 45,700 years ago with the population ancestral to both New Guineans and Oceanians. Such a late date for the New Guinean group could indicate Denisovan survival as late as 14,500 years ago, which would make them the latest surviving archaic human species. A third wave appears to have introgressed into East Asia, but there is not enough DNA evidence to pinpoint a solid timeframe.
The mtDNA from Denisova 4 bore a high similarity to that of Denisova 3, indicating that they belonged to the same population. The genetic diversity among the Denisovans from Denisova Cave is on the lower range of what is seen in modern humans, and is comparable to that of Neanderthals. However, it is possible that the inhabitants of Denisova Cave were more or less reproductively isolated from other Denisovans, and that, across their entire range, Denisovan genetic diversity may have been much higher.
Denisova Cave, over time of habitation, continually swung from a fairly warm and moderately humid pine and birch forest to tundra or forest-tundra landscape. Conversely, Baishiya Karst Cave is situated at a high elevation, an area characterized by low temperature, low oxygen, and poor resource availability. Colonization of high-altitude regions, due to such harsh conditions, was previously assumed to have only been accomplished by modern humans. Denisovans seem to have also inhabited the jungles of Southeast Asia. The Tam Ngu Hao 2 site might have been a closed forest environment.
Anatomy
Little is known of the precise anatomical features of the Denisovans since the only physical remains discovered so far are a finger bone, four teeth, long bone fragments, a partial jawbone, a parietal bone skull fragment, and a rib bone. The finger bone is within the modern human range of variation for women, which is in contrast to the large, robust molars which are more similar to those of Middle to Late Pleistocene archaic humans. The third molar is outside the range of any Homo species except H. habilis and H. rudolfensis, and is more like those of australopithecines. The second molar is larger than those of modern humans and Neanderthals, and is more similar to those of H. erectus and H. habilis. Like Neanderthals, the mandible had a gap behind the molars, and the front teeth were flattened; but Denisovans lacked a high mandibular body, and the mandibular symphysis at the midline of the jaw was more receding. The parietal is reminiscent of that of H. erectus.
A facial reconstruction has been generated by comparing methylation at individual genetic loci associated with facial structure. This analysis suggested that Denisovans, much like Neanderthals, had a long, broad, and projecting face; large nose; sloping forehead; protruding jaw; elongated and flattened skull; and wide chest and hips. The Denisovan tooth row was longer than that of Neanderthals and anatomically modern humans.
Middle-to-Late Pleistocene East Asian archaic human skullcaps typically share features with Neanderthals. The skullcaps from Xuchang feature prominent brow ridges like Neanderthals, though the nuchal and angular tori near the base of the skull are either reduced or absent, and the back of the skull is rounded off like in early modern humans. Xuchang 1 had a large brain volume of approximately 1800 cc, on the high end for Neanderthals and early modern humans, and well beyond the present-day human average.
The Denisovan genome from Denisova Cave has variants of genes which, in modern humans, are associated with dark skin, brown hair, and brown eyes. The Denisovan genome also contains a variant region around the EPAS1 gene that in Tibetans assists with adaptation to low oxygen levels at high elevation, and in a region containing the WARS2 and TBX15 loci which affect body-fat distribution in the Inuit. In Papuans, introgressed Neanderthal alleles are highest in frequency in genes expressed in the brain, whereas Denisovan alleles have highest frequency in genes expressed in bones and other tissue.
Culture
Denisova Cave

 Some ornaments (above) and animal bones and stone tools (below) found in Denisova Cave. Note, ornaments may have been crafted by modern humans
Some ornaments (above) and animal bones and stone tools (below) found in Denisova Cave. Note, ornaments may have been crafted by modern humans
Early Middle Paleolithic stone tools from Denisova Cave included cores, scrapers, denticulate tools, and notched tools, deposited about 287±41 thousand years ago in the Main Chamber of the cave; and about 269±97 thousand years ago in the South Chamber; up to 170±19 thousand and 187±14 thousand years ago in the Main and East Chambers, respectively.
Middle Paleolithic assemblages were dominated by flat, discoidal, and Levallois cores, and there were some isolated sub-prismatic cores. There were predominantly side scrapers (a scraper with only the sides used to scrape), but also notched-denticulate tools, end-scrapers (a scraper with only the ends used to scrape), burins, chisel-like tools, and truncated flakes. These dated to 156±15 thousand years ago in the Main Chamber, 58±6 thousand years ago in the East Chamber, and 136±26–47±8 thousand years ago in the South Chamber.
Early Upper Paleolithic artefacts date to 44±5 thousand years ago in the Main Chamber, 63±6 thousand years ago in the East Chamber, and 47±8 thousand years ago in the South Chamber, though some layers of the East Chamber seem to have been disturbed. There was blade production and Levallois production, but scrapers were again predominant. A well-developed, Upper Paleolithic stone bladelet technology distinct from the previous scrapers began accumulating in the Main Chamber around 36±4 thousand years ago.
In the Upper Paleolithic layers, there were also several bone tools and ornaments: a marble ring, an ivory ring, an ivory pendant, a red deer tooth pendant, an elk tooth pendant, a chloritolite bracelet, and a bone needle. However, Denisovans are only confirmed to have inhabited the cave until 55 ka; the dating of Upper Paleolithic artefacts overlaps with modern human migration into Siberia (though there are no occurrences in the Altai region); and the DNA of the only specimen in the cave dating to the time interval (Denisova 14) is too degraded to confirm species identity, so the attribution of these artefacts is unclear.
Tibet

The inhabitants of Baishiya Karst Cave seem to have been extensively processing goat antelopes, cows, deer, horses, and woolly rhinoceros. They were also butchering large carnivores (cave hyena, dog, and big cat), marmots, hare, and eagles. They may have also used these animals' long bones to make bone tools, and additionally there are stone artefacts in each layer excavated.
In 1998, five child hand- and footprint impressions were discovered in a travertine unit near the Quesang hot springs in Tibet; in 2021, they were dated to 226 to 169 thousand years ago using uranium decay dating. This is the oldest evidence of human occupation of the Tibetan Plateau, and since the Xiahe mandible is the oldest human fossil from the region (though younger than the Quesang impressions), these may have been made by Denisovan children. The impressions were printed onto a small panel of space, and there is little overlap between all the prints, so they seem to have been taking care to make new imprints in unused space. If considered art, they are the oldest known examples of rock art. Similar hand stencils and impressions do not appear again in the archeological record until roughly 40,000 years ago.
The footprints comprise four right impressions and one left superimposed on one of the rights. They were probably left by two individuals. The tracks of the individual who superimposed their left onto their right may have scrunched up their toes and wiggled them in the mud, or dug their finger into the toe prints. The footprints average 192.3 mm (7.57 in) long, which roughly equates to a 7 or 8 year old child by modern human growth rates. There are two sets of handprints (from a left and right hand), which may have been created by an older child unless one of the former two individuals had long fingers. The handprints average 161.1 mm (6.34 in), which roughly equates with a 12 year old modern human child, and the middle finger length agrees with a 17 year old modern human. One of the handprints shows an impression of the forearm, and the individual was wiggling their thumb through the mud.
Interbreeding
See also: Interbreeding between archaic and modern humansAnalyses of the modern human genomes indicate past interbreeding with at least two groups of archaic humans, Neanderthals and Denisovans, and that such interbreeding events occurred on multiple occasions. Comparisons of the Denisovan, Neanderthal, and modern human genomes have revealed evidence of a complex web of interbreeding among these lineages.
Archaic humans
As much as 17% of the Denisovan genome from Denisova Cave represents DNA from the local Neanderthal population. Denisova 11 was an F1 (first generation) Denisovan/Neanderthal hybrid; the fact that such an individual was found may indicate interbreeding was a common occurrence here. The Denisovan genome shares more derived alleles with the Altai Neanderthal genome from Siberia than with the Vindija Cave Neanderthal genome from Croatia or the Mezmaiskaya cave Neanderthal genome from the Caucasus, suggesting that the gene flow came from a population that was more closely related to the local Altai Neanderthals. However, Denny's Denisovan father had the typical Altai Neanderthal introgression, while her Neanderthal mother represented a population more closely related to Vindija Neanderthals. Denisova 25, dated to 200ka, is estimated to have inherited 5% of his genome from a previously unknown population of Neanderthals, and came from a different population of Denisovans than the younger samples.
About 4% of the Denisovan genome derives from an unidentified archaic hominin, perhaps the source of the anomalous ancient mtDNA, indicating this species diverged from Neanderthals and humans over a million years ago. The only identified Homo species of Late Pleistocene Asia are H. erectus and H. heidelbergensis, though in 2021, specimens allocated to the latter species were reclassified as H. longi and H. daliensis.
Before splitting from Neanderthals, their ancestors ("Neandersovans") migrating into Europe apparently interbred with an unidentified "superarchaic" human species who were already present there; these superarchaics were the descendants of a very early migration out of Africa around 1.9 mya.
Modern humans
A 2011 study found that Denisovan DNA is prevalent in Papuans, Aboriginal Australians, Near Oceanians, Polynesians, Fijians, Eastern Indonesians, and Aeta (from the Philippines); but not in East Asians, western Indonesians, Jahai people (from Malaysia), or Onge (from the Andaman Islands). This may suggest that Denisovan introgression occurred within the Pacific region rather than on the Asian mainland, and that ancestors of the latter groups were not present in Southeast Asia at the time. In the Melanesian genome, about 4–6% or 1.9–3.4% derives from Denisovan introgression. Prior to 2021, New Guineans and Australian Aborigines were reported to have the most introgressed DNA, but Australians have less than New Guineans. A 2021 study discovered 30 to 40% more Denisovan ancestry in Aeta people in the Philippines than in Papuans, estimated as about 5% of the genome. The Aeta Magbukon in Luzon have the highest known proportion of Denisovan ancestry of any population in the world. In Papuans, less Denisovan ancestry is seen in the X chromosome than autosomes, and some autosomes (such as chromosome 11) also have less Denisovan ancestry, which could indicate hybrid incompatibility. The former observation could also be explained by less female Denisovan introgression into modern humans, or more female modern human immigrants who diluted Denisovan X chromosome ancestry.
In contrast, 0.2% derives from Denisovan ancestry in mainland Asians and Native Americans. South Asians were found to have levels of Denisovan admixture similar to that seen in East Asians. The discovery of the 40,000-year-old Chinese modern human Tianyuan Man lacking Denisovan DNA significantly different from the levels in modern-day East Asians discounts the hypothesis that immigrating modern humans simply diluted Denisovan ancestry whereas Melanesians lived in reproductive isolation. A 2018 study of Han Chinese, Japanese, and Dai genomes showed that modern East Asians have DNA from two different Denisovan populations: one similar to the Denisovan DNA found in Papuan genomes, and a second that is closer to the Denisovan genome from Denisova Cave. This could indicate two separate introgression events involving two different Denisovan populations. In South Asian genomes, DNA only came from the same single Denisovan introgression seen in Papuans. A 2019 study found a third wave of Denisovans which introgressed into East Asians. Introgression, also, may not have immediately occurred when modern humans immigrated into the region.
The timing of introgression into Oceanian populations likely occurred after Eurasians and Oceanians split roughly 58,000 years ago, and before Papuan and Aboriginal Australians split from each other roughly 37,000 years ago. Given the present day distribution of Denisovan DNA, this may have taken place in Wallacea, though the discovery of a 7,200 year old Toalean girl (closely related to Papuans and Aboriginal Australians) from Sulawesi carrying Denisovan DNA makes Sundaland another potential candidate. Other early Sunda hunter gatherers so far sequenced carry very little Denisovan DNA, which either means the introgression event did not take place in Sundaland, or Denisovan ancestry was diluted with gene flow from the mainland Asian Hòabìnhian culture and subsequent Neolithic cultures.
In other regions of the world, archaic introgression into humans stems from a group of Neanderthals related to those which inhabited Vindija Cave, Croatia, as opposed to archaics related to Siberian Neanderthals and Denisovans. However, about 3.3% of the archaic DNA in the modern Icelandic genome descends from the Denisovans, and such a high percentage could indicate a western Eurasian population of Denisovans which introgressed into either Vindija-related Neanderthals or immigrating modern humans.
Denisovan genes may have helped early modern humans migrating out of Africa to acclimatize. Although not present in the sequenced Denisovan genome, the distribution pattern and divergence of HLA-B*73 from other HLA alleles (involved in the immune system's natural killer cell receptors) has led to the suggestion that it introgressed from Denisovans into modern humans in West Asia. In a 2011 study, half of the HLA alleles of modern Eurasians were shown to represent archaic HLA haplotypes, and were inferred to be of Denisovan or Neanderthal origin. A haplotype of EPAS1 in modern Tibetans, which allows them to live at high elevations in a low-oxygen environment, likely came from Denisovans. Genes related to phospholipid transporters (which are involved in fat metabolism) and to trace amine-associated receptors (involved in smelling) are more active in people with more Denisovan ancestry. Denisovan genes may have conferred a degree of immunity against the G614 mutation of SARS-CoV-2. Denisovan introgressions may have influenced the immune system of present-day Papuans and potentially favoured "variants to immune-related phenotypes" and "adaptation to the local environment".
In December 2023, scientists reported that genes inherited by modern humans from Neanderthals and Denisovans may biologically influence the daily routine of modern humans.
See also
- Red Deer Cave people – Prehistoric humans from 12,500 BCE in southwest China
- Homo longi – Archaic human from China, 146,000 BP
- Homo naledi – South African archaic human species
- Nesher Ramla Homo – Extinct population of archaic humans
- Timeline of human evolution
References
- Zimmer, Carl (2 March 2024). "On the Trail of the Denisovans - DNA has shown that the extinct humans thrived around the world, from chilly Siberia to high-altitude Tibet — perhaps even in the Pacific islands". The New York Times. Archived from the original on 2 March 2024. Retrieved 2 March 2024.
- ^ Krause, J.; Fu, Q.; Good, J. M.; Viola, B.; et al. (2010). "The complete mitochondrial DNA genome of an unknown hominin from southern Siberia". Nature. 464 (7290): 894–897. Bibcode:2010Natur.464..894K. doi:10.1038/nature08976. ISSN 1476-4687. PMC 10152974. PMID 20336068. S2CID 4415601.
- Douglas, M. M.; Douglas, J. M. (2016). Exploring Human Biology in the Laboratory. Morton Publishing Company. p. 324. ISBN 9781617313905. Archived from the original on 1 September 2024. Retrieved 25 October 2020.
- Zubova, A.; Chikisheva, T.; Shunkov, M. V. (2017). "The Morphology of Permanent Molars from the Paleolithic Layers of Denisova Cave". Archaeology, Ethnology & Anthropology of Eurasia. 45: 121–134. doi:10.17746/1563-0110.2017.45.1.121-134.
- McKie, Robin (30 March 2024). "Scientists link elusive human group to 150,000-year-old Chinese 'dragon man'". The Observer. ISSN 0029-7712. Archived from the original on 1 September 2024. Retrieved 31 March 2024.
- ^ Bae, C. J. (2024). "The "Muddle in the Middle" (~400 ka–~100 ka)". The Paleoanthropology of Eastern Asia. University of Hawai‘i Press. pp. 95–131. doi:10.1515/9780824898106-007. ISBN 9780824898106.
- Ovodov, N. D.; Crockford, S. J.; Kuzmin, Y. V.; Higham, T. F.; et al. (2011). "A 33,000-Year-Old Incipient Dog from the Altai Mountains of Siberia: Evidence of the Earliest Domestication Disrupted by the Last Glacial Maximum". PLOS ONE. 6 (7): e22821. Bibcode:2011PLoSO...622821O. doi:10.1371/journal.pone.0022821. PMC 3145761. PMID 21829526.
- Reich, D. (2018). Who We Are and How We Got Here. Oxford University Press. p. 53. ISBN 978-0-19-882125-0.
- ^ Douka, K. (2019). "Age estimates for hominin fossils and the onset of the Upper Palaeolithic at Denisova Cave". Nature. 565 (7741): 640–644. Bibcode:2019Natur.565..640D. doi:10.1038/s41586-018-0870-z. PMID 30700871. S2CID 59525455. Archived from the original on 6 May 2020. Retrieved 7 December 2019.
- ^ Jacobs, Zenobia; Li, Bo; Shunkov, Michael V.; Kozlikin, Maxim B.; et al. (January 2019). "Timing of archaic hominin occupation of Denisova Cave in southern Siberia". Nature. 565 (7741): 594–599. Bibcode:2019Natur.565..594J. doi:10.1038/s41586-018-0843-2. ISSN 1476-4687. PMID 30700870. S2CID 59525956. Archived from the original on 7 May 2020. Retrieved 29 May 2020.
- ^ Slon, V.; Viola, B.; Renaud, G.; Gansauge, M.-T.; et al. (2017). "A fourth Denisovan individual". Science Advances. 3 (7): e1700186. Bibcode:2017SciA....3E0186S. doi:10.1126/sciadv.1700186. PMC 5501502. PMID 28695206.
- ^ Gibbons, Ann (11 July 2024). "The most ancient human genome yet has been sequenced—and it's a Denisovan's". Science. doi:10.1126/science.zi9n4zp. Archived from the original on 18 July 2024. Retrieved 13 July 2024.
- ^ Sawyer, S.; Renaud, G.; Viola, B.; Hublin, J.-J.; et al. (2015). "Nuclear and mitochondrial DNA sequences from two Denisovan individuals". Proceedings of the National Academy of Sciences. 112 (51): 15696–700. Bibcode:2015PNAS..11215696S. doi:10.1073/pnas.1519905112. PMC 4697428. PMID 26630009.
- ^ Bennett, E. A.; Crevecoeur, I.; Viola, B.; et al. (2019). "Morphology of the Denisovan phalanx closer to modern humans than to Neanderthals". Science Advances. 5 (9): eaaw3950. Bibcode:2019SciA....5.3950B. doi:10.1126/sciadv.aaw3950. PMC 6726440. PMID 31517046.
- ^ Gibbons, Anne (2019). "First fossil jaw of Denisovans finally puts a face on elusive human relatives". Science. doi:10.1126/science.aax8845. S2CID 188493848.
- ^ Chen, F.; Welker, F.; Shen, C.-C.; et al. (2019). "A late Middle Pleistocene Denisovan mandible from the Tibetan Plateau" (PDF). Nature. 569 (7756): 409–412. Bibcode:2019Natur.569..409C. doi:10.1038/s41586-019-1139-x. PMID 31043746. S2CID 141503768. Archived (PDF) from the original on 13 December 2019. Retrieved 7 December 2019.
- Shang, D.; et al. (2020). "Denisovan DNA in Late Pleistocene sediments from Baishiya Karst Cave on the Tibetan Plateau". Science. 370 (6516): 584–587. doi:10.1126/science.abb6320. PMID 33122381. S2CID 225956074.
- ^ Xia, Huan; Zhang, Dongju; Wang, Jian; Fagernäs, Zandra; Li, Ting; Li, Yuanxin; Yao, Juanting; Lin, Dongpeng; Troché, Gaudry; Smith, Geoff M.; Chen, Xiaoshan; Cheng, Ting; Shen, Xuke; Han, Yuanyuan; Olsen, Jesper V. (3 July 2024). "Middle and Late Pleistocene Denisovan subsistence at Baishiya Karst Cave". Nature. 632 (8023): 108–113. doi:10.1038/s41586-024-07612-9. ISSN 1476-4687. PMC 11291277. PMID 38961285.
- ^ Demeter, F.; Zanolli, C.; Westaway, K. E.; et al. (2022). "A Middle Pleistocene Denisovan molar from the Annamite Chain of northern Laos". Nature Communications. 13 (2557): 2557. Bibcode:2022NatCo..13.2557D. doi:10.1038/s41467-022-29923-z. PMC 9114389. PMID 35581187.
- ^ Callaway, Ewen (2010). "Fossil genome reveals ancestral link". Nature. 468 (7327): 1012. Bibcode:2010Natur.468.1012C. doi:10.1038/4681012a. PMID 21179140.
- Ao, H.; Liu, C.-R.; Roberts, A. P. (2017). "An updated age for the Xujiayao hominin from the Nihewan Basin, North China: Implications for Middle Pleistocene human evolution in East Asia". Journal of Human Evolution. 106: 54–65. Bibcode:2017AGUFMPP13B1080A. doi:10.1016/j.jhevol.2017.01.014. hdl:1885/232536. PMID 28434540.
- ^ Cooper, A.; Stringer, C. B. (2013). "Did the Denisovans Cross Wallace's Line?". Science. 342 (6156): 321–23. Bibcode:2013Sci...342..321C. doi:10.1126/science.1244869. PMID 24136958. S2CID 206551893.
- ^ Warren, M. (2019). "Biggest Denisovan fossil yet spills ancient human's secrets". Nature News. 569 (7754): 16–17. Bibcode:2019Natur.569...16W. doi:10.1038/d41586-019-01395-0. PMID 31043736.
- Ji, Qiang; Wu, Wensheng; Ji, Yannan; Li, Qiang; et al. (25 June 2021). "Late Middle Pleistocene Harbin cranium represents a new Homo species". The Innovation. 2 (3): 100132. Bibcode:2021Innov...200132J. doi:10.1016/j.xinn.2021.100132. ISSN 2666-6758. PMC 8454552. PMID 34557772.
- ^ Reich, D.; Green, R. E.; Kircher, M.; et al. (2010). "Genetic history of an archaic hominin group from Denisova Cave in Siberia" (PDF). Nature. 468 (7327): 1053–60. Bibcode:2010Natur.468.1053R. doi:10.1038/nature09710. hdl:10230/25596. PMC 4306417. PMID 21179161. Archived from the original on 17 May 2020. Retrieved 29 July 2018.
- Brown, S.; Higham, T.; Slon, V.; Pääbo, S. (2016). "Identification of a new hominin bone from Denisova Cave, Siberia using collagen fingerprinting and mitochondrial DNA analysis". Scientific Reports. 6: 23559. Bibcode:2016NatSR...623559B. doi:10.1038/srep23559. PMC 4810434. PMID 27020421.
- ^ Viola, B. T.; Gunz, P.; Neubauer, S. (2019). "A parietal fragment from Denisova cave". 88th Annual Meeting of the American Association of Physical Anthropologists. Archived from the original on 26 September 2019. Retrieved 18 January 2020.
- ^ Lao, O.; Bertranpetit, J.; Mondal, M. (2019). "Approximate Bayesian computation with deep learning supports a third archaic introgression in Asia and Oceania". Nature Communications. 10 (1): 246. Bibcode:2019NatCo..10..246M. doi:10.1038/s41467-018-08089-7. ISSN 2041-1723. PMC 6335398. PMID 30651539.
- Rogers, A. R.; Bohlender, R. J.; Huff, C. D. (2017). "Early history of Neanderthals and Denisovans". Proceedings of the National Academy of Sciences. 114 (37): 9859–9863. Bibcode:2017PNAS..114.9859R. doi:10.1073/pnas.1706426114. PMC 5604018. PMID 28784789.
- Ho, K. K. (2016). "Hominin interbreeding and the evolution of human variation". Journal of Biological Research-Thessaloniki. 23: 17. doi:10.1186/s40709-016-0054-7. PMC 4947341. PMID 27429943.
- Malyarchuk, B. A. (2011). "Adaptive evolution of the Homo mitochondrial genome". Molecular Biology. 45 (5): 845–850. doi:10.1134/S0026893311050104. PMID 22393781. S2CID 43284294.
- Pääbo, S.; Kelso, J.; Reich, D.; Slatkin, M.; et al. (2014). "The complete genome sequence of a Neanderthal from the Altai Mountains". Nature. 505 (7481): 43–49. Bibcode:2014Natur.505...43P. doi:10.1038/nature12886. ISSN 1476-4687. PMC 4031459. PMID 24352235.
- Kuhlwilm, M.; Gronau, I.; Hubisz, M. J.; de Filippo, C.; et al. (2016). "Ancient gene flow from early modern humans into Eastern Neanderthals". Nature. 530 (7591): 429–433. Bibcode:2016Natur.530..429K. doi:10.1038/nature16544. ISSN 1476-4687. PMC 4933530. PMID 26886800.
- Posth, C.; Wißing, C.; Kitagawa, K.; Pagani, L.; et al. (2017). "Deeply divergent archaic mitochondrial genome provides lower time boundary for African gene flow into Neanderthals". Nature Communications. 8: 16046. Bibcode:2017NatCo...816046P. doi:10.1038/ncomms16046. ISSN 2041-1723. PMC 5500885. PMID 28675384.
- Bertranpetit, J.; Majumder, P. P.; Li, Q.; Laayouni, H.; et al. (2016). "Genomic analysis of Andamanese provides insights into ancient human migration into Asia and adaptation". Nature Genetics. 48 (9): 1066–1070. doi:10.1038/ng.3621. hdl:10230/34401. ISSN 1546-1718. PMID 27455350. S2CID 205352099.
- Callaway, E. (2013). "Hominin DNA baffles experts". Nature. 504 (7478): 16–17. Bibcode:2013Natur.504...16C. doi:10.1038/504016a. PMID 24305130.
- Tattersall, I. (2015). The Strange Case of the Rickety Cossack and other Cautionary Tales from Human Evolution. Palgrave Macmillan. p. 200. ISBN 978-1-137-27889-0.
- Meyer, M.; Arsuaga, J.-L.; et al. (2016). "Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins". Nature. 531 (7595): 504–507. Bibcode:2016Natur.531..504M. doi:10.1038/nature17405. PMID 26976447. S2CID 4467094.
- Callaway, E. (2011). "First Aboriginal genome sequenced". Nature News. doi:10.1038/news.2011.551.
- ^ Reich, David; Patterson, Nick; Kircher, Martin; Delfin, Frederick; et al. (2011). "Denisova Admixture and the First Modern Human Dispersals into Southeast Asia and Oceania". The American Journal of Human Genetics. 89 (4): 516–28. doi:10.1016/j.ajhg.2011.09.005. PMC 3188841. PMID 21944045.
- ^ Skov, L.; Macià, M. C.; Sveinbjörnsson, G.; et al. (2020). "The nature of Neanderthal introgression revealed by 27,566 Icelandic genomes". Nature. 582 (7810): 78–83. Bibcode:2020Natur.582...78S. doi:10.1038/s41586-020-2225-9. PMID 32494067. S2CID 216076889.
- ^ Jacobs, G. S.; Hudjashov, G.; Saag, L.; Kusuma, P.; et al. (2019). "Multiple Deeply Divergent Denisovan Ancestries in Papuans". Cell. 177 (4): 1010–1021.e32. doi:10.1016/j.cell.2019.02.035. hdl:1983/7df38af7-d075-4444-9111-b859650f6d38. ISSN 0092-8674. PMID 30981557.
- "Denisovan DNA may help modern humans adapt to different environments". Archived from the original on 11 August 2024. Retrieved 11 August 2024.
- ^ Larena, Maximilian; McKenna, James; Sanchez-Quinto, Federico; Bernhardsson, Carolina; et al. (August 2021). "Philippine Ayta possess the highest level of Denisovan ancestry in the world". Current Biology. 31 (19): 4219–4230.e10. Bibcode:2021CBio...31E4219L. doi:10.1016/j.cub.2021.07.022. PMC 8596304. PMID 34388371. S2CID 236994320.
- Callaway, E. (2019). "Siberia's ancient ghost clan starts to surrender its secrets". Nature News. 566 (7745): 444–446. Bibcode:2019Natur.566..444C. doi:10.1038/d41586-019-00672-2. PMID 30814723.
- Gokhman, D.; Lavi, E.; Prüfer, K.; Fraga, M. F.; et al. (2014). "Reconstructing the DNA methylation maps of the Neandertal and the Denisovan". Science. 344 (6183): 523–27. Bibcode:2014Sci...344..523G. doi:10.1126/science.1250368. PMID 24786081. S2CID 28665590.
- Gokhman, D.; Mishol, N.; de Manuel, M.; Marques-Bonet, T.; et al. (2019). "Reconstructing Denisovan Anatomy Using DNA Methylation Maps". Cell. 179 (1): 180–192. doi:10.1016/j.cell.2019.08.035. PMID 31539495. S2CID 202676502.
- Li, Z.-Y.; Wu, X.-J.; Zhou, L.-.P; et al. (2017). "Late Pleistocene archaic human crania from Xuchang, China". Science. 355 (6328): 969–972. Bibcode:2017Sci...355..969L. doi:10.1126/science.aal2482. PMID 28254945. S2CID 206654741.
- ^ Meyer, M.; Kircher, M.; Gansauge, M.-T.; et al. (2012). "A High-Coverage Genome Sequence from an Archaic Denisovan Individual". Science. 338 (6104): 222–226. Bibcode:2012Sci...338..222M. doi:10.1126/science.1224344. PMC 3617501. PMID 22936568.
- ^ Huerta-Sánchez, E.; Jin, X.; et al. (2014). "Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA". Nature. 512 (7513): 194–97. Bibcode:2014Natur.512..194H. doi:10.1038/nature13408. PMC 4134395. PMID 25043035.
- Racimo, Fernando; Gokhman, David; Fumagalli, Matteo; Ko, Amy; et al. (2017). "Archaic Adaptive Introgression in TBX15/WARS2". Molecular Biology and Evolution. 34 (3): 509–524. doi:10.1093/molbev/msw283. PMC 5430617. PMID 28007980.
- Akkuratov, Evgeny E.; Gelfand, Mikhail S.; Khrameeva, Ekaterina E. (2018). "Neanderthal and Denisovan ancestry in Papuans: A functional study". Journal of Bioinformatics and Computational Biology. 16 (2): 1840011. doi:10.1142/S0219720018400115. PMID 29739306.
- Dennell, R. (2019). "Dating of hominin discoveries at Denisova". Nature News. 565 (7741): 571–572. Bibcode:2019Natur.565..571D. doi:10.1038/d41586-019-00264-0. PMID 30700881.
- ^ Zhang, D. D.; Bennett, M. R.; Cheng, H.; Wang, L.; et al. (2021). "Earliest parietal art: Hominin hand and foot traces from the middle Pleistocene of Tibet". Science Bulletin. 66 (24): 2506–2515. Bibcode:2021SciBu..66.2506Z. doi:10.1016/j.scib.2021.09.001. ISSN 2095-9273. PMID 36654210. S2CID 239102132.
- ^ Pennisi, E. (2013). "More Genomes from Denisova Cave Show Mixing of Early Human Groups". Science. 340 (6134): 799. Bibcode:2013Sci...340..799P. doi:10.1126/science.340.6134.799. PMID 23687020.
- Green RE, Krause J, Briggs AW, et al. (2010). "A draft sequence of the Neandertal genome" (PDF). Science. 328 (5979): 710–22. Bibcode:2010Sci...328..710G. doi:10.1126/science.1188021. PMC 5100745. PMID 20448178. Archived (PDF) from the original on 13 August 2012. Retrieved 3 May 2013.
- Warren, M. (2018). "Mum's a Neanderthal, Dad's a Denisovan: First discovery of an ancient-human hybrid – Genetic analysis uncovers a direct descendant of two different groups of early humans". Nature. 560 (7719): 417–418. Bibcode:2018Natur.560..417W. doi:10.1038/d41586-018-06004-0. PMID 30135540.
- ^ Prüfer, K.; Racimo, Fernando; Patterson, N.; Jay, F.; Sankararaman, S.; Sawyer, S.; et al. (2013). "The complete genome sequence of a Neanderthal from the Altai Mountains". Nature. 505 (7481): 43–49. Bibcode:2014Natur.505...43P. doi:10.1038/nature12886. PMC 4031459. PMID 24352235.
- Warren, Matthew (2018). "Mum's a Neanderthal, Dad's a Denisovan: First discovery of an ancient-human hybrid". Nature. 560 (7719): 417–18. Bibcode:2018Natur.560..417W. doi:10.1038/d41586-018-06004-0. PMID 30135540.
- Wolf, A. B.; Akey, J. M. (2018). "Outstanding questions in the study of archaic hominin admixture". PLOS Genetics. 14 (5): e1007349. doi:10.1371/journal.pgen.1007349. PMC 5978786. PMID 29852022.
- Ni, X.; Ji, Q.; Wu, W.; et al. (2021). "Massive cranium from Harbin in northeastern China establishes a new Middle Pleistocene human lineage". Innovation. 2 (3): 100130. Bibcode:2021Innov...200130N. doi:10.1016/j.xinn.2021.100130. ISSN 2666-6758. PMC 8454562. PMID 34557770. S2CID 236784246.
- Rogers, A. R.; Harris, N. S.; Achenbach, A. A. (2020). "Neanderthal-Denisovan ancestors interbred with a distantly related hominin". Science Advances. 6 (8): eaay5483. Bibcode:2020SciA....6.5483R. doi:10.1126/sciadv.aay5483. PMC 7032934. PMID 32128408.
- Yang, Melinda A. (6 January 2022). "A genetic history of migration, diversification, and admixture in Asia". Human Population Genetics and Genomics. 2 (1): 1–32. doi:10.47248/hpgg2202010001. ISSN 2770-5005. Archived from the original on 16 April 2023. Retrieved 5 December 2023.
- Vernot, B.; et al. (2016). "Excavating Neandertal and Denisovan DNA from the genomes of Melanesian individuals". Science. 352 (6282): 235–239. Bibcode:2016Sci...352..235V. doi:10.1126/science.aad9416. PMC 6743480. PMID 26989198.
- Rasmussen, M.; Guo, Xiaosen; Wang, Yong; Lohmueller, Kirk E.; Rasmussen, Simon; et al. (2011). "An Aboriginal Australian genome reveals separate human dispersals into Asia". Science. 334 (6052): 94–98. Bibcode:2011Sci...334...94R. doi:10.1126/science.1211177. PMC 3991479. PMID 21940856.
- Prüfer, K.; Racimo, F.; Patterson, N.; Jay, F.; et al. (2013). "The complete genome sequence of a Neanderthal from the Altai Mountains". Nature. 505 (7481): 43–49. Bibcode:2014Natur.505...43P. doi:10.1038/nature12886. PMC 4031459. PMID 24352235.
- ^ Browning, S. R.; Browning, B. L.; Zhou, Yi.; Tucci, S.; et al. (2018). "Analysis of Human Sequence Data Reveals Two Pulses of Archaic Denisovan Admixture". Cell. 173 (1): 53–61.e9. doi:10.1016/j.cell.2018.02.031. ISSN 0092-8674. PMC 5866234. PMID 29551270.
- Fu, Q.; Meyer, M.; Gao, X.; et al. (2013). "DNA analysis of an early modern human from Tianyuan Cave, China". Proceedings of the National Academy of Sciences. 110 (6): 2223–2227. Bibcode:2013PNAS..110.2223F. doi:10.1073/pnas.1221359110. PMC 3568306. PMID 23341637.
- Carlhoff, Selina (2021). "Genome of a middle Holocene hunter-gatherer from Wallace". Nature. 596 (7873): 543–547. Bibcode:2021Natur.596..543C. doi:10.1038/s41586-021-03823-6. PMC 8387238. PMID 34433944.
- Abi-Rached, L.; Jobin, M. J.; Kulkarni, S.; McWhinnie, A.; et al. (2011). "The Shaping of Modern Human Immune Systems by Multiregional Admixture with Archaic Humans". Science. 334 (6052): 89–94. Bibcode:2011Sci...334...89A. doi:10.1126/science.1209202. PMC 3677943. PMID 21868630.
- Sankararaman, S.; Mallick, S.; Patterson, N.; Reich, D. (2016). "The combined landscape of Denisovan and Neanderthal ancestry in present-day humans". Current Biology. 26 (9): 1241–1247. Bibcode:2016CBio...26.1241S. doi:10.1016/j.cub.2016.03.037. PMC 4864120. PMID 27032491.
- Muscat, Baron Y. (2021). "Could the Denisovan Genes have conferred enhanced Immunity Against the G614 Mutation of SARS-CoV-2?". Human Evolution. 36. Archived (PDF) from the original on 19 July 2021. Retrieved 19 July 2021.
- Vespasiani, Davide M.; Jacobs, Guy S.; Cook, Laura E.; Brucato, Nicolas; Leavesley, Matthew; Kinipi, Christopher; Ricaut, François-Xavier; Cox, Murray P.; Gallego Romero, Irene (8 December 2022). "Denisovan introgression has shaped the immune system of present-day Papuans". PLOS Genetics. 18 (12): e1010470. doi:10.1371/journal.pgen.1010470. ISSN 1553-7390. PMC 9731433. PMID 36480515.
- Zimmer, Carl (14 December 2023). "Morning Person? You Might Have Neanderthal Genes to Thank. - Hundreds of genetic variants carried by Neanderthals and Denisovans are shared by people who like to get up early". The New York Times. Archived from the original on 14 December 2023. Retrieved 14 December 2023.
Further reading
- Karlsson, Mattis. From Fossil To Fact: The Denisova Discovery as Science in Action (Thesis). LiU E-press. ISBN 9789179291716. Retrieved 18 March 2022.
External links
 Media related to Denisova at Wikimedia Commons
Media related to Denisova at Wikimedia Commons- The Denisova Consortium's raw sequence data and alignments
- Human Timeline (Interactive) – Smithsonian, National Museum of Natural History (August 2016).
- Picture of Denisovan molar.
| Human evolution | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Taxonomy (Hominins) |
| ||||||||||||||||||||||||||
| Ancestors |
| ||||||||||||||||||||||||||
| Models |
| ||||||||||||||||||||||||||
| Timelines | |||||||||||||||||||||||||||
| Others | |||||||||||||||||||||||||||
| Prehistoric technology | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||
| |||||||||||||||||||||
| |||||||||||||||||||||