| Revision as of 14:00, 7 May 2007 view sourceTonyrenploki (talk | contribs)30 edits →Overview of biological functions← Previous edit | Revision as of 14:01, 7 May 2007 view source Tonyrenploki (talk | contribs)30 edits →Interactions with proteinsNext edit → | ||
| Line 13: | Line 13: | ||
| Tonyrenploki | Tonyrenploki | ||
| ==Interactions with proteins== | |||
| All the functions of DNA depend on interactions with proteins. These protein interactions can be non-specific, or the protein can bind specifically to a single DNA sequence. Enzymes can also bind to DNA and of these, the polymerases that copy the DNA base sequence in transcription and DNA replication are particularly important. | |||
| ===DNA-binding proteins=== | |||
| <div class="thumb tleft" style="background-color: #f9f9f9; border: 1px solid #CCCCCC; margin:0.5em;"> | |||
| {|border="0" width=260px border="0" cellpadding="0" cellspacing="0" style="font-size: 85%; border: 1px solid #CCCCCC; margin: 0.3em;" | |||
| |] | |||
| |- | |||
| |] | |||
| |} | |||
| <div style="border: none; width:260px;"><div class="thumbcaption">Interaction of DNA with ]s (shown in white, top). These proteins' basic amino acids (below left, blue) bind to the acidic phosphate groups on DNA (below right, red).</div></div></div> | |||
| Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called ]. In eukaryotes this structure involves DNA binding to a complex of small basic proteins called ]s, while in prokaryotes multiple types of proteins are involved.<ref>{{cite journal | author = Sandman K, Pereira S, Reeve J | title = Diversity of prokaryotic chromosomal proteins and the origin of the nucleosome | journal = Cell Mol Life Sci | volume = 54 | issue = 12 | pages = 1350 – 64 | year = 1998 | id = PMID 9893710}}</ref><ref>{{cite journal |author=Dame RT |title=The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin |journal=Mol. Microbiol. |volume=56 |issue=4 |pages=858-70 |year=2005 |pmid=15853876}}</ref> The histones form a disk-shaped complex called a ], which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones making ]s to the acidic sugar-phosphate backbone of the DNA, and are therefore largely independent of the base sequence.<ref>{{cite journal | author = Luger K, Mäder A, Richmond R, Sargent D, Richmond T | title = Crystal structure of the nucleosome core particle at 2.8 A resolution | journal = Nature | volume = 389 | issue = 6648 | pages = 251 – 60 | year = 1997 | id = PMID 9305837}}</ref> Chemical modifications of these basic amino acid residues include ], ] and ].<ref>{{cite journal | author = Jenuwein T, Allis C | title = Translating the histone code | journal = Science | volume = 293 | issue = 5532 | pages = 1074 – 80 | year = 2001 | id = PMID 11498575}}</ref> These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to ]s and changing the rate of transcription.<ref>{{cite journal | author = Ito T | title = Nucleosome assembly and remodelling | journal = Curr Top Microbiol Immunol | volume = 274 | issue = | pages = 1 – 22 | year = | id = PMID 12596902}}</ref> Other non-specific DNA-binding proteins found in chromatin include the high-mobility group proteins, which bind preferentially to bent or distorted DNA.<ref>{{cite journal | author = Thomas J | title = HMG1 and 2: architectural DNA-binding proteins | journal = Biochem Soc Trans | volume = 29 | issue = Pt 4 | pages = 395 – 401 | year = 2001 | id = PMID 11497996}}</ref> These proteins are important in bending arrays of nucleosomes and arranging them into more complex chromatin structures.<ref>{{cite journal | author = Grosschedl R, Giese K, Pagel J | title = HMG domain proteins: architectural elements in the assembly of nucleoprotein structures | journal = Trends Genet | volume = 10 | issue = 3 | pages = 94-100 | year = 1994 | id = PMID 8178371}}</ref> | |||
| A distinct group of DNA-binding proteins are the single-stranded-DNA-binding proteins that specifically bind single-stranded DNA. In humans, replication protein A is the best-characterised member of this family and is essential for most processes where the double helix is separated, including DNA replication, recombination and DNA repair.<ref>{{cite journal | author = Iftode C, Daniely Y, Borowiec J | title = Replication protein A (RPA): the eukaryotic SSB | journal = Crit Rev Biochem Mol Biol | volume = 34 | issue = 3 | pages = 141 – 80 | year = 1999 | id = PMID 10473346}}</ref> These binding proteins seem to stabilize single-stranded DNA and protect it from forming ]s or being degraded by ]s. | |||
| ] transcription factor bound to its DNA target<ref>Created from </ref>]] | |||
| In contrast, other proteins have evolved to specifically bind particular DNA sequences. The most intensively studied of these are the various classes of ]s, which are proteins that regulate transcription. Each one of these proteins bind to one particular set of DNA sequences and thereby activates or inhibits the transcription of genes with these sequences close to their ]s. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins; this locates the polymerase at the promoter and allows it to begin transcription.<ref>{{cite journal | author = Myers L, Kornberg R | title = Mediator of transcriptional regulation | journal = Annu Rev Biochem | volume = 69 | issue = | pages = 729 – 49 | year = | id = PMID 10966474}}</ref> Alternatively, transcription factors can bind ]s that modify the histones at the promoter; this will change the accessibility of the DNA template to the polymerase.<ref>{{cite journal | author = Spiegelman B, Heinrich R | title = Biological control through regulated transcriptional coactivators | journal = Cell | volume = 119 | issue = 2 | pages = 157-67 | year = 2004 | id = PMID 15479634}}</ref> | |||
| As these DNA targets can occur throughout an organism's genome, changes in the activity of one type of transcription factor can affect thousands of genes.<ref>{{cite journal | author = Li Z, Van Calcar S, Qu C, Cavenee W, Zhang M, Ren B | title = A global transcriptional regulatory role for c-Myc in Burkitt's lymphoma cells | url=http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pubmed&pubmedid=12808131 | journal = Proc Natl Acad Sci U S A | volume = 100 | issue = 14 | pages = 8164 – 9 | year = 2003 | id = PMID 12808131}}</ref> Consequently, these proteins are often the targets of the ] processes that mediate responses to environmental changes or cellular differentiation and development. The specificity of these transcription factors' interactions with DNA come from the proteins making multiple contacts to the edges of the DNA bases, allowing them to "read" the DNA sequence. Most of these base-interactions are made in the major groove, where the bases are most accessible.<ref>{{cite journal | author = Pabo C, Sauer R | title = Protein-DNA recognition | journal = Annu Rev Biochem | volume = 53 | issue = | pages = 293 – 321 | year = | id = PMID 6236744}}</ref> | |||
| ] ] (green) in a complex with its substrate DNA<ref>Created from </ref>]] | |||
| ===DNA-modifying enzymes=== | |||
| ====Nucleases and ligases==== | |||
| Nucleases are ]s that cut DNA strands by catalyzing the ] of the ]s. Nucleases that hydrolyse nucleotides from the ends of DNA strands are called ]s, while ]s cut within strands. The most frequently-used nucleases in ] are the ], which cut DNA at specific sequences. For instance, the EcoRV enzyme shown to the left recognizes the 6-base sequence 5′-GAT|ATC-3′ and makes a cut at the vertical line. In nature, these enzymes protect ] against ] infection by digesting the phage DNA when it enters the bacterial cell, acting as part of the ].<ref>{{cite journal | author = Bickle T, Krüger D | title = Biology of DNA restriction | url=http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=372918&blobtype=pdf | journal = Microbiol Rev | volume = 57 | issue = 2 | pages = 434 – 50 | year = 1993 | id = PMID 8336674}}</ref> In technology, these sequence-specific nucleases are used in ] and ]. | |||
| Enzymes called ]s can rejoin cut or broken DNA strands, using the energy from either ] or ].<ref name=Doherty>{{cite journal | author = Doherty A, Suh S | title = Structural and mechanistic conservation in DNA ligases. | url=http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pubmed&pubmedid=11058099 | journal = Nucleic Acids Res | volume = 28 | issue = 21 | pages = 4051 – 8 | year = 2000 | id = PMID 11058099}}</ref> Ligases are particularly important in ] DNA replication, as they join together the short segments of DNA produced at the ] into a complete copy of the DNA template. They are also used in ] and ].<ref name=Doherty/> | |||
| ====Topoisomerases and helicases==== | |||
| ]s are enzymes with both nuclease and ligase activity. These proteins change the amount of ] in DNA. Some of these enzyme work by cutting the DNA helix and allowing one section to rotate, thereby reducing its level of supercoiling; the enzyme then seals the DNA break.<ref name=Champoux/> Other types of these enzymes are capable of cutting one DNA helix and then passing a second strand of DNA through this break, before rejoining the helix.<ref>{{cite journal | author = Schoeffler A, Berger J | title = Recent advances in understanding structure-function relationships in the type II topoisomerase mechanism | journal = Biochem Soc Trans | volume = 33 | issue = Pt 6 | pages = 1465 – 70 | year = 2005 | id = PMID 16246147}}</ref> Topoisomerases are required for many processes involving DNA, such as DNA replication and transcription.<ref name=Wang/> | |||
| ]s are proteins that are a type of ]. They use the chemical energy in ]s, predominantly ], to break hydrogen bonds between bases and unwind the DNA double helix into single strands.<ref>{{cite journal | author = Tuteja N, Tuteja R | title = Unraveling DNA helicases. Motif, structure, mechanism and function | url=http://www.blackwell-synergy.com/links/doi/10.1111%2Fj.1432-1033.2004.04094.x | journal = Eur J Biochem | volume = 271 | issue = 10 | pages = 1849-63 | year = 2004 | id = PMID 15128295}}</ref> These enzymes are essential for most processes where enzymes need to access the DNA bases. | |||
| ====Polymerases==== | |||
| Polymerases are enzymes that synthesise polynucleotide chains from ]s. They function by adding nucleotides onto the 3′ ] of the previous nucleotide in the DNA strand. As a consequence, all polymerases work in a 5′ to 3′ direction.<ref name=Joyce>{{cite journal | author = Joyce C, Steitz T | title = Polymerase structures and function: variations on a theme? | url=http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=177480&blobtype=pdf | journal = J Bacteriol | volume = 177 | issue = 22 | pages = 6321 – 9 | year = 1995 | id = PMID 7592405}}</ref> In the ] of these enzymes, the nucleoside triphosphate substrate base-pairs to a single-stranded polynucleotide template: this allows polymerases to accurately synthesise the complementary strand of this template. Polymerases are classified according to the type of template that they use. | |||
| In ], a DNA-dependent ] makes a DNA copy of a DNA sequence. Accuracy is vital in this process, so many of these polymerases have a ] activity. Here, the polymerase recognizes the occasional mistakes in the synthesis reaction by the lack of base pairing between the mismatched nucleotides. If a mismatch is detected, a 3′ to 5′ ] activity is activated and the incorrect base removed.<ref>{{cite journal | author = Hubscher U, Maga G, Spadari S | title = Eukaryotic DNA polymerases | journal = Annu Rev Biochem | volume = 71 | issue = | pages = 133 – 63 | year = | id = PMID 12045093}}</ref> In most organisms DNA polymerases function in a large complex called the ] that contains multiple accessory subunits, such as the ] or ]s.<ref>{{cite journal | author = Johnson A, O'Donnell M | title = Cellular DNA replicases: components and dynamics at the replication fork | journal = Annu Rev Biochem | volume = 74 | issue = | pages = 283 – 315 | year = | id = PMID 15952889}}</ref> | |||
| RNA-dependent DNA polymerases are a specialised class of polymerases that copy the sequence of an RNA strand into DNA. They include ], which is a ] enzyme involved in the infection of cells by ]es, and ], which is required for the replication of ]s.<ref>{{cite journal | author = Tarrago-Litvak L, Andréola M, Nevinsky G, Sarih-Cottin L, Litvak S | title = The reverse transcriptase of HIV-1: from enzymology to therapeutic intervention | url=http://www.fasebj.org/cgi/reprint/8/8/497 | journal = FASEB J | volume = 8 | issue = 8 | pages = 497-503 | year = 1994 | id = PMID 7514143}}</ref><ref name=Greider/> Telomerase is an unusual polymerase because it contains its own RNA template as part of its structure.<ref name=Nugent/> | |||
| Transcription is carried out by a DNA-dependent ] that copies the sequence of a DNA strand into RNA. To begin transcribing a gene, the RNA polymerase binds to a sequence of DNA called a ] and separates the DNA strands. It then copies the gene sequence into a ] transcript until it reaches a region of DNA called the ], where it halts and detaches from the DNA. As with human DNA-dependent DNA polymerases, RNA polymerase II, the enzyme that transcribes most of the genes in the human genome, operates as part of a large protein complex with multiple regulatory and accessory subunits.<ref>{{cite journal | author = Martinez E | title = Multi-protein complexes in eukaryotic gene transcription | journal = Plant Mol Biol | volume = 50 | issue = 6 | pages = 925 – 47 | year = 2002 | id = PMID 12516863}}</ref> | |||
| ==Genetic recombination== | ==Genetic recombination== | ||
Revision as of 14:01, 7 May 2007
For other uses, see DNA (disambiguation).
Deoxyribonucleic acid, or DNA is a nucleic acid molecule that contains the genetic instructions used in the development and functioning of all living organisms. The main role of DNA is the long-term storage of information and it is often compared to a set of blueprints, since DNA contains the instructions needed to construct other components of cells, such as proteins and RNA molecules. The DNA segments that carry this genetic information are called genes, but other DNA sequences have structural purposes, or are involved in regulating the use of this genetic information.
Chemically, DNA is a long polymer of simple units called nucleotides, which are held together by a backbone made of alternating sugars and phosphate groups. Attached to each sugar is one of four types of molecules called bases. It is the sequence of these four bases along the backbone that encodes information. This information is read using the genetic code, which specifies the sequence of the amino acids within proteins. The code is read by copying stretches of DNA into the related nucleic acid RNA, in a process called transcription. Most of these RNA molecules are used to synthesize proteins, but others are used directly in structures such as ribosomes and spliceosomes.
Within cells, DNA is organized into structures called chromosomes and the set of chromosomes within a cell make up a genome. These chromosomes are duplicated before cells divide, in a process called DNA replication. Eukaryotic organisms such as animals, plants, and fungi store their DNA inside the cell nucleus, while in prokaryotes such as bacteria it is found in the cell's cytoplasm. Within the chromosomes, chromatin proteins such as histones compact and organize DNA, which helps control its interactions with other proteins and thereby control which genes are transcribed.
Tonyrenploki
Genetic recombination

|

|
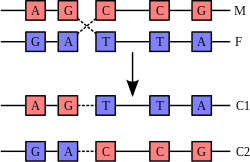
A DNA helix does not usually interact with other segments of DNA, and in human cells the different chromosomes even occupy separate areas in the nucleus called "chromosome territories". This physical separation of different chromosomes is important for the ability of DNA to function as a stable repository for information, as one of the few times chromosomes interact is when they recombine. Recombination is when two DNA helices break, swap a section and then rejoin.
Recombination allows chromosomes to exchange genetic information and produces new combinations of genes, which increases the efficiency of natural selection and can be important in the rapid evolution of new proteins. Genetic recombination can also be involved in DNA repair, particularly in the cell's response to double-strand breaks.
The most common form of recombination is homologous recombination, where the two chromosomes involved share very similar sequences. Non-homologous recombination can be damaging to cells, as it can produce chromosomal translocations and genetic abnormalities. The recombination reaction is catalyzed by enzymes known as recombinases, such as Cre recombinase. In the first step, the recombinase creates a nick in one strand of a DNA double helix, allowing the nicked strand to pull apart from its complementary strand and anneal to one strand of the double helix on the opposite chromatid. A second nick allows the strand in the second chromatid to pull apart and anneal to the remaining strand in the first helix, forming a structure known as a cross-strand exchange or a Holliday junction. The Holliday junction is a tetrahedral junction structure that can be moved along the pair of chromosomes, swapping one strand for another. The recombination reaction is then halted by cleavage of the junction and re-ligation of the released DNA.
Evolution of DNA-based metabolism
DNA contains the genetic information that allows all modern living things to function, grow and reproduce. However, it is unclear how long in the 4-billion-year history of life DNA has performed this function, as it has been proposed that the earliest forms of life may have used RNA as their genetic material. RNA may have acted as the central part of early cell metabolism as it can both transmit genetic information and carry out catalysis as part of ribozymes. This ancient RNA world where nucleic acid would have been used for both catalysis and genetics may have influenced the evolution of the current genetic code based on four nucleotide bases. This would occur since the number of unique bases in such an organism is a trade-off between a small number of bases increasing replication accuracy and a large number of bases increasing the catalytic efficiency of ribozymes.
Unfortunately, there is no direct evidence of ancient genetic systems, as recovery of DNA from most fossils is impossible. This is because DNA will survive in the environment for less than one million years and slowly degrades into short fragments in solution. Although claims for older DNA have been made, most notably a report of the isolation of a viable bacterium from a salt crystal 250-million years old, these claims are controversial and have been disputed.
Uses in technology
Further information: ]Modern biology and biochemistry make intensive use of recombinant DNA technology. Recombinant DNA is a man-made DNA sequence assembled from other DNA sequences in a plasmid. These plasmids can be transformed into organisms. The genetically modified organisms produced can be used to produce products such as recombinant proteins, used in medical research, or be grown in agriculture.
Forensics
Further information: ]Forensic scientists can use DNA in blood, semen, skin, saliva or hair at a crime scene to identify a perpetrator. This process is called genetic fingerprinting, or more accurately, DNA profiling. In DNA profiling, the lengths of variable sections of repetitive DNA, such as short tandem repeats and minisatellites, are compared between people. This method is usually an extremely reliable technique for identifying a criminal. However, identification can be complicated if the scene is contaminated with DNA from several people. DNA profiling was developed in 1984 by British geneticist Sir Alec Jeffreys, and first used in forensic science to convict Colin Pitchfork in the 1988 Enderby murders case. People convicted of certain types of crimes may be required to provide a sample of DNA for a database. This has helped investigators solve old cases where only a DNA sample was obtained from the scene. DNA profiling can also be used to identify victims of mass casualty incidents.
Bioinformatics
Further information: ]Bioinformatics involves the manipulation, searching, and data mining of DNA sequence data. The development of techniques to store and search DNA sequences have led to widely-applied advances in computer science, especially string searching algorithms, machine learning and database theory. String searching or matching algorithms, which find an occurrence of a sequence of letters inside a larger sequence of letters, were developed to search for specific sequences of nucleotides. In other applications such as text editors, even simple algorithms for this problem usually suffice, but DNA sequences cause these algorithms to exhibit near-worst-case behaviour due to their small number of distinct characters. The related problem of sequence alignment aims to identify homologous sequences and locate the specific mutations that make them distinct. These techniques, especially multiple sequence alignment, are used in studying phylogenetic relationships and protein function. Data sets representing entire genomes' worth of DNA sequences, such as those produced by the Human Genome Project, are difficult to use without annotations, which label the locations of genes and regulatory elements on each chromosome. Regions of DNA sequence that have the characteristic patterns associated with protein- or RNA-coding genes can be identified by gene finding algorithms, which allow researchers to predict the presence of particular gene products in an organism even before they have been isolated experimentally.
DNA and computation
Further information: ]DNA was first used in computing to solve a small version of the directed Hamiltonian path problem, an NP-complete problem. DNA computing is advantageous over electronic computers in power use, space use, and efficiency, due to its ability to compute in a highly parallel fashion (see parallel computing). A number of other problems, including simulation of various abstract machines, the boolean satisfiability problem, and the bounded version of the travelling salesman problem, have since been analysed using DNA computing. Due to its compactness, DNA also has a theoretical role in cryptography, where in particular it allows unbreakable one-time pads to be efficiently constructed and used.
History and anthropology
Further information: ]Because DNA collects mutations over time, which are then inherited, it contains historical information and by comparing DNA sequences, geneticists can infer the evolutionary history of organisms, their phylogeny. This field of phylogenetics is a powerful tool in evolutionary biology. If DNA sequences within a species are compared, population geneticists can learn the history of particular populations. This can be used in studies ranging from ecological genetics to anthropology; for example, DNA evidence is being used to try to identify the Ten Lost Tribes of Israel.
DNA has also been used to look at modern family relationships, such as establishing family relationships between the descendants of Sally Hemings and Thomas Jefferson. This usage is closely related to the use of DNA in criminal investigations detailed above. Indeed, some criminal investigations have been solved when DNA from crime scenes has matched relatives of the guilty individual.
History

DNA was first isolated by Friedrich Miescher who, in 1869, discovered a microscopic substance in the pus of discarded surgical bandages. As it resided in the nuclei of cells, he called it "nuclein". In 1929 this discovery was followed by Phoebus Levene's identification of the base, sugar and phosphate nucleotide unit. Levene suggested that DNA consisted of a string of nucleotide units linked together through the phosphate groups. However, Levene thought the chain was short and the bases repeated in a fixed order. In 1937 William Astbury produced the first X-ray diffraction patterns that showed that DNA had a regular structure.
In 1943, Oswald Theodore Avery discovered that traits of the "smooth" form of the Pneumococcus could be transferred to the "rough" form of the same bacteria by mixing killed "smooth" bacteria with the live "rough" form. Avery identified DNA as this transforming principle. DNA's role in heredity was confirmed in 1953, when Alfred Hershey and Martha Chase in the Hershey-Chase experiment showed that DNA is the genetic material of the T2 phage.
In 1953, based on X-ray diffraction images taken by Rosalind Franklin and the information that the bases were paired, James D. Watson and Francis Crick suggested what is now accepted as the first accurate model of DNA structure in the journal Nature. Experimental evidence for Watson and Crick's model were published in a series of five articles in the same issue of Nature. Of these, Franklin and Raymond Gosling's paper saw the publication of the X-ray diffraction image, which was key in Watson and Crick interpretation, as well as another article, co-authored by Maurice Wilkins and his colleagues. Franklin and Gosling's subsequent paper identified the distinctions between the A and B structures of the double helix in DNA. In 1962 Watson, Crick, and Maurice Wilkins jointly received the Nobel Prize in Physiology or Medicine (Franklin didn't share the prize with them since she had died earlier).
In an influential presentation in 1957, Crick laid out the "Central Dogma" of molecular biology, which foretold the relationship between DNA, RNA, and proteins, and articulated the "adaptor hypothesis". Final confirmation of the replication mechanism that was implied by the double-helical structure followed in 1958 through the Meselson-Stahl experiment. Further work by Crick and coworkers showed that the genetic code was based on non-overlapping triplets of bases, called codons, allowing Har Gobind Khorana, Robert W. Holley and Marshall Warren Nirenberg to decipher the genetic code. These findings represent the birth of molecular biology.
See also
- Genetic disorder
- Plasmid
- DNA sequencing
- Southern blot
- DNA microarray
- Polymerase chain reaction
- Protein-DNA interaction site predictor
- Phosphoramidite
- Quantification of nucleic acids
References
- Created from PDB 1M6G
- Cremer T, Cremer C (2001). "Chromosome territories, nuclear architecture and gene regulation in mammalian cells". Nat Rev Genet. 2 (4): 292–301. PMID 11283701.
- Pál C, Papp B, Lercher M (2006). "An integrated view of protein evolution". Nat Rev Genet. 7 (5): 337–48. PMID 16619049.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - O'Driscoll M, Jeggo P (2006). "The role of double-strand break repair - insights from human genetics". Nat Rev Genet. 7 (1): 45–54. PMID 16369571.
- Ghosh K, Van Duyne G (2002). "Cre-loxP biochemistry". Methods. 28 (3): 374–83. PMID 12431441.
- Dickman M, Ingleston S, Sedelnikova S, Rafferty J, Lloyd R, Grasby J, Hornby D (2002). "The RuvABC resolvasome". Eur J Biochem. 269 (22): 5492–501. PMID 12423347.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Joyce G (2002). "The antiquity of RNA-based evolution". Nature. 418 (6894): 214–21. PMID 12110897.
- Orgel L. "Prebiotic chemistry and the origin of the RNA world" (PDF). Crit Rev Biochem Mol Biol. 39 (2): 99–123. PMID 15217990.
- Davenport R (2001). "Ribozymes. Making copies in the RNA world". Science. 292 (5520): 1278. PMID 11360970.
- Szathmáry E (1992). "What is the optimum size for the genetic alphabet?" (PDF). Proc Natl Acad Sci U S A. 89 (7): 2614–8. PMID 1372984.
- Lindahl T (1993). "Instability and decay of the primary structure of DNA". Nature. 362 (6422): 709–15. PMID 8469282.
- Vreeland R, Rosenzweig W, Powers D (2000). "Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal". Nature. 407 (6806): 897–900. PMID 11057666.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Hebsgaard M, Phillips M, Willerslev E (2005). "Geologically ancient DNA: fact or artefact?". Trends Microbiol. 13 (5): 212–20. PMID 15866038.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Nickle D, Learn G, Rain M, Mullins J, Mittler J (2002). "Curiously modern DNA for a "250 million-year-old" bacterium". J Mol Evol. 54 (1): 134–7. PMID 11734907.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Houdebine L. "Transgenic animal models in biomedical research". Methods Mol Biol. 360: 163–202. PMID 17172731.
- Daniell H, Dhingra A (2002). "Multigene engineering: dawn of an exciting new era in biotechnology". Curr Opin Biotechnol. 13 (2): 136–41. PMID 11950565.
- Job D (2002). "Plant biotechnology in agriculture". Biochimie. 84 (11): 1105–10. PMID 12595138.
- Collins A, Morton N (1994). "Likelihood ratios for DNA identification" (PDF). Proc Natl Acad Sci U S A. 91 (13): 6007–11. PMID 8016106.
- Weir B, Triggs C, Starling L, Stowell L, Walsh K, Buckleton J (1997). "Interpreting DNA mixtures". J Forensic Sci. 42 (2): 213–22. PMID 9068179.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Jeffreys A, Wilson V, Thein S. "Individual-specific 'fingerprints' of human DNA". Nature. 316 (6023): 76–9. PMID 2989708.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Colin Pitchfork — first murder conviction on DNA evidence also clears the prime suspect Forensic Science Service Accessed 23 Dec 2006
- "DNA Identification in Mass Fatality Incidents". National Institute of Justice. September 2006.
- Baldi, Pierre. Brunak, Soren. Bioinformatics: The Machine Learning Approach MIT Press (2001) ISBN 978-0-262-02506-5
- Gusfield, Dan. Algorithms on Strings, Trees, and Sequences: Computer Science and Computational Biology. Cambridge University Press, 15 January 1997. ISBN 978-0-521-58519-4.
- Sjölander K (2004). "Phylogenomic inference of protein molecular function: advances and challenges". Bioinformatics. 20 (2): 170–9. PMID 14734307.
- Mount DM (2004). Bioinformatics: Sequence and Genome Analysis (2 ed.). Cold Spring Harbor Laboratory Press. ISBN 0879697121.
{{cite book}}: Text "Cold Spring Harbor, NY" ignored (help); Text "location" ignored (help) - Adleman L (1994). "Molecular computation of solutions to combinatorial problems". Science. 266 (5187): 1021–4. PMID 7973651.
- Parker J (2003). "Computing with DNA". EMBO Rep. 4 (1): 7–10. PMID 12524509.
- Ashish Gehani, Thomas LaBean and John Reif. DNA-Based Cryptography. Proceedings of the 5th DIMACS Workshop on DNA Based Computers, Cambridge, MA, USA, 14 – 15 June 1999.
- Wray G (2002). "Dating branches on the tree of life using DNA". Genome Biol. 3 (1): REVIEWS0001. PMID 11806830.
- Lost Tribes of Israel, NOVA, PBS airdate: 22 February 2000. Transcript available from PBS.org, (last accessed on 4 March 2006)
- Kleiman, Yaakov. "The Cohanim/DNA Connection: The fascinating story of how DNA studies confirm an ancient biblical tradition". aish.com (January 13, 2000). Accessed 4 March 2006.
- Bhattacharya, Shaoni. "Killer convicted thanks to relative's DNA". newscientist.com (20 April 2004). Accessed 22 Dec 06
- Dahm R (2005). "Friedrich Miescher and the discovery of DNA". Dev Biol. 278 (2): 274–88. PMID 15680349.
- Levene P, (1919). "The structure of yeast nucleic acid". J Biol Chem. 40 (2): 415–24.
{{cite journal}}: CS1 maint: extra punctuation (link) - Astbury W, (1947). "Nucleic acid". Symp. SOC. Exp. Bbl. 1 (66).
{{cite journal}}: CS1 maint: extra punctuation (link) - Avery O, MacLeod C, McCarty M (1944). "Studies on the chemical nature of the substance inducing transformation of pneumococcal types. Inductions of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus type III". J Exp Med. 79 (2): 137–158.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Hershey A, Chase M (1952). "Independent functions of viral protein and nucleic acid in growth of bacteriophage" (PDF). J Gen Physiol. 36 (1): 39–56. PMID 12981234.
- ^ Watson J.D. and Crick F.H.C. "A Structure for Deoxyribose Nucleic Acid". (PDF) Nature 171, 737 – 738 (1953). Accessed 13 Feb 2007.
- Cite error: The named reference
Watsonwas invoked but never defined (see the help page). - Nature Archives Double Helix of DNA: 50 Years
- Molecular Configuration in Sodium Thymonucleate. Franklin R. and Gosling R.G.Nature 171, 740 – 741 (1953)Nature Archives Full Text (PDF)
- Molecular Structure of Deoxypentose Nucleic Acids. Wilkins M.H.F., A.R. Stokes A.R. & Wilson, H.R. Nature 171, 738 – 740 (1953)Nature Archives (PDF)
- Evidence for 2-Chain Helix in Crystalline Structure of Sodium Deoxyribonucleate. Franklin R. and Gosling R.G. Nature 172, 156 – 157 (1953)Nature Archives,full text (PDF)
- The Nobel Prize in Physiology or Medicine 1962 Nobelprize .org Accessed 22 Dec 06
- Crick, F.H.C. On degenerate templates and the adaptor hypothesis (PDF). genome.wellcome.ac.uk (Lecture, 1955). Accessed 22 Dec 2006
- Meselson M, Stahl F (1958). "The replication of DNA in Escherichia coli". Proc Natl Acad Sci U S A. 44 (7): 671–82. PMID 16590258.
- The Nobel Prize in Physiology or Medicine 1968 Nobelprize.org Accessed 22 Dec 06
Further reading
- Clayton, Julie. (Ed.). 50 Years of DNA, Palgrave MacMillan Press, 2003. ISBN 978-1-40-391479-8
- Judson, Horace Freeland. The Eighth Day of Creation: Makers of the Revolution in Biology, Cold Spring Harbor Laboratory Press, 1996. ISBN 978-0-87-969478-4
- Olby, Robert. The Path to The Double Helix: Discovery of DNA, first published in October 1974 by MacMillan, with foreword by Francis Crick; ISBN 978-0-48-668117-7; the definitive DNA textbook, revised in 1994, with a 9 page postscript.
- Ridley, Matt. Francis Crick: Discoverer of the Genetic Code (Eminent Lives) HarperCollins Publishers; 192 pp, ISBN 978-0-06-082333-7 2006
- Rose, Steven. The Chemistry of Life, Penguin, ISBN 978-0-14-027273-4.
- Watson, James D. and Francis H.C. Crick. A structure for Deoxyribose Nucleic Acid (PDF). Nature 171, 737 – 738, 25 April 1953.
- Watson, James D. DNA: The Secret of Life ISBN 978-0-375-41546-3.
- Watson, James D. The Double Helix: A Personal Account of the Discovery of the Structure of DNA (Norton Critical Editions). ISBN 978-0-393-95075-5
- Watson, James D. "Avoid boring people and other lessons from a life in science" New York: Random House. ISBN 978-0-375-421844 (0-375-41284-0)366pp 2007
- Calladine, Chris R.; Drew, Horace R.; Luisi, Ben F. and Travers, Andrew A. Understanding DNA, Elsevier Academic Press, 2003. ISBN 978-0-12155089-9
DVD
- DNA — The Story of the Pioneers who Changed the World, Windfall Films Production for Channel Four Television & PBS Thirteen-WNET — 2003, PAL , NTSC PBS Shop
- DNA interactive PAL , NTSC ,
- DNA: The Secret of Life Carolina Biological
- DNA — Secret of Photo 51 Rosalind Franklin — NOVA documentary (NTSC — Region 1?)
- Cracking the Code of Life NOVA documentary (NTSC — All Regions)
External links
Listen to this article(2 parts, 34 minutes)
- Crick's personal papers at Mandeville Special Collections Library, Geisel Library, University of California, San Diego
- DNA Interactive (requires Adobe Flash)
- DNA from the beginning
- Double Helix 1953 – 2003 National Centre for Biotechnology Education
- Double helix: 50 years of DNA, Nature
- Rosalind Franklin's contributions to the study of DNA
- U.S. National DNA Day — watch videos and participate in real-time chat with top scientists
- Genetic Education Modules for Teachers — DNA from the Beginning Study Guide
- Listen to Francis Crick and James Watson talking on the BBC in 1962, 1972, and 1974
- PDB Molecule of the Month DNA
- DNA under electron microscope
- Template:Dmoz
- DNA Articles — articles and information collected from various sources
- Template:McGrawHillAnimation
- DNA coiling to form chromosomes
- DISPLAR: DNA binding site prediction on protein
- Dolan DNA Learning Center
- Olby, R. (2003) "Quiet debut for the double helix" Nature 421 (January 23): 402 – 405.
- Basic animated guide to DNA cloning
- DNA the Double Helix Game From the official Nobel Prize web site
| Types of nucleic acids | |||||||
|---|---|---|---|---|---|---|---|
| Constituents | |||||||
| Ribonucleic acids (coding, non-coding) |
| ||||||
| Deoxyribonucleic acids | |||||||
| Analogues | |||||||
| Cloning vectors | |||||||
Template:Link FA Template:Link FA
Categories: